- Review: found
Review of 'The global distribution of Crimean-Congo hemorrhagic fever'
Average rating: | Rated 2.5 of 5. |
Level of importance: | Rated 2 of 5. |
Level of validity: | Rated 2 of 5. |
Level of completeness: | Rated 2 of 5. |
Level of comprehensibility: | Rated 4 of 5. |
Competing interests: | None |
Reviewed article
- Record: found
- Abstract: found
- Article: found
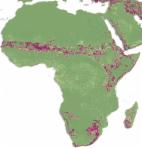
The global distribution of Crimean-Congo hemorrhagic fever
Review information
Review text
This paper attempts to produce a SDM (species distribution model) for the distribution of Crimean-Congo hemorrhagic fever virus, which is an importan pathogens of humans. The authors used a large dataset of human clinical cases to map the distribution of the clinical cases (not the virus) and they attempt to correlate with several envionmental variables to map the expected geographical distribution of the virus.
First, I would like to kindly call the attention of the authors about the methods. Using a (partial) distribution of human clinical cases is not enough to map where the virus exist. Literally dozens of reports exist about the silent foci of the virus. These silent foci are "places" where the critical threshold of infected ticks is not large enough as to produce observable clinical cases. But the virus is still there. I acknowledge that it is very hard to obtain a comprehensive view of the distribution of CCHF virus (providing that it is a BSL-4 organism and that PCR is the best procedure -expensive- to map such distribution). However, the compiled distribution of clinical cases is also partial: it miss literally thousands of records from countries like Turkey (which, by the way, are publicly available).
Stating it clearly (and humbly): the map of clinical cases *does not* correspond with the distribution of the virus. I honestly believe that introducing the dataset of human cases, introduces an extra layer of "noise", because the human habits and contact with ticks, the presence of reservoirs, the distribution of the involved ticks, etc., are nor included in the basic dataset.
Second, I would like to focus on the explanatory variables. It is well know that gridded datasets of climate are not adequate to map the "habitat suitability" of an organism. They tend to be biased because interpolation procedures. It has been demonstrated several times that satellite images are much better for these purpose. Again, I must to acknowledge that processing satellite images is time-comsuming (but you have also publicly available and processed datasets...). Since the authors used *both* gridded data and satellite imagery, then a problem of interpolation and integration arises (see several papers by Peterson, Lobo, Araujo, etc.). Further on this, it is well known (actually, is a paradigm) that rainfall has no effect on ticks (the only known vectors of the virus) but relative humidity or saturation deficit have a clear effect. "Warm and arid" zones cannot be catched with "rainfall" but with reative humidity.
I must to agree that authors want to model "CHHF occurrence", which still has not a meaning in my mind: CCHF presence, CCHF human cases? In any case, my modest point(s) of view are:
1. authors shoud avoid to use human cases as a "proxy" for virus persistence: it depends upon vertebrates and ticks, and then human contcat with the infected areas
2. environmental variables are poorly chosen, and they will not reflect the behaviour of the tick vectors
3. there are not covariates explaining the distribution of the vertebrates that could feed the ticks
4. there are not covariates explaingin the "critical" step of human behaviours and contcat with foci of ticks.
5. the dataset of human cases is partial: as a result, you can easily seen that the distribution of the virus does not correlate with the actual known prevalence of the virus in ticks...
That said, I would like to kindly propose the authors to update the dataset of clinical cases, to choose variables that adequately explain the behaviour of the tick and the vertebrates and then update the distribution maps. Statistical methods are correct, in my modest point of view, and they should be the ground or future estimations of this important thrad to human health.