- Record: found
- Abstract: found
- Article: found
Coevolution of URAT1 and Uricase during Primate Evolution: Implications for Serum Urate Homeostasis and Gout
Read this article at
Abstract
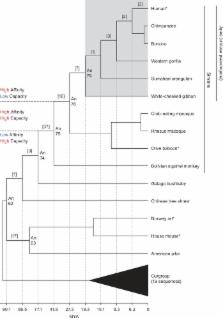
Uric acid is the highly insoluble end-product of purine metabolism in humans. Serum levels exceeding the solubility threshold can trigger formation of urate crystals resulting in gouty arthritis. Uric acid is primarily excreted through the kidneys with 90% reabsorbed back into the bloodstream through the uric acid transporter URAT1. This reabsorption process is essential for the high serum uric acid levels found in humans. We discovered that URAT1 proteins from humans and baboons have higher affinity for uric acid compared with transporters from rats and mice. This difference in transport kinetics of URAT1 orthologs, along with inability of modern apes to oxidize uric acid due to loss of the uricase enzyme, prompted us to ask whether these events occurred concomitantly during primate evolution. Ancestral URAT1 sequences were computationally inferred and ancient transporters were resurrected and assayed, revealing that affinity for uric acid was increased during the evolution of primates. This molecular fine-tuning occurred between the origins of simians and their diversification into New- and Old-World monkey and ape lineages. Remarkably, it was driven in large-part by only a few amino acid replacements within the transporter. This alteration in primate URAT1 coincided with changes in uricase that greatly diminished the enzymatic activity and took place 27–77 Ma. These results suggest that the modifications to URAT1 transporters were potentially adaptive and that maintaining more constant, high levels of serum uric acid may have provided an advantage to our primate ancestors.
Related collections
Most cited references19
- Record: found
- Abstract: found
- Article: not found
Codon-substitution models for heterogeneous selection pressure at amino acid sites.

- Record: found
- Abstract: found
- Article: found
Tree of Life Reveals Clock-Like Speciation and Diversification
- Record: found
- Abstract: found
- Article: not found