- Record: found
- Abstract: found
- Article: found
Parsing interindividual drug variability: an emerging role for systems pharmacology
Read this article at
Abstract
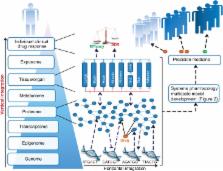
There is notable interindividual heterogeneity in drug response, affecting both drug efficacy and toxicity, resulting in patient harm and the inefficient utilization of limited healthcare resources. Pharmacogenomics is at the forefront of research to understand interindividual drug response variability, but although many genotype-drug response associations have been identified, translation of pharmacogenomic associations into clinical practice has been hampered by inconsistent findings and inadequate predictive values. These limitations are in part due to the complex interplay between drug-specific, human body and environmental factors influencing drug response and therefore pharmacogenomics, whilst intrinsically necessary, is by itself unlikely to adequately parse drug variability. The emergent, interdisciplinary and rapidly developing field of systems pharmacology, which incorporates but goes beyond pharmacogenomics, holds significant potential to further parse interindividual drug variability. Systems pharmacology broadly encompasses two distinct research efforts, pharmacologically-orientated systems biology and pharmacometrics. Pharmacologically-orientated systems biology utilizes high throughput omics technologies, including next-generation sequencing, transcriptomics and proteomics, to identify factors associated with differential drug response within the different levels of biological organization in the hierarchical human body. Increasingly complex pharmacometric models are being developed that quantitatively integrate factors associated with drug response. Although distinct, these research areas complement one another and continual development can be facilitated by iterating between dynamic experimental and computational findings. Ultimately, quantitative data-derived models of sufficient detail will be required to help realize the goal of precision medicine. WIREs Syst Biol Med 2015, 7:221–241. doi: 10.1002/wsbm.1302
Related collections
Most cited references69
- Record: found
- Abstract: found
- Article: not found
A randomized trial of genotype-guided dosing of warfarin.
- Record: found
- Abstract: found
- Article: not found
Immune self-reactivity triggered by drug-modified HLA-peptide repertoire.
- Record: found
- Abstract: found
- Article: not found