- Record: found
- Abstract: found
- Article: found
Comparative FISH mapping of ribosomal DNA clusters and TTAGG telomeric sequences to holokinetic chromosomes of eight species of the insect order Psocoptera

Abstract
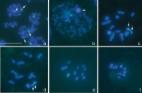
Repetitive DNAs are the main components of eukaryotic genome. We mapped the 18S rDNA and TTAGG telomeric probe sequences by FISH to meiotic chromosomes of eight species of the order Psocoptera considered a basal taxon of Paraneoptera : Valenzuela burmeisteri (Brauer, 1876), Stenopsocus lachlani Kolbe, 1960, Graphopsocus cruciatus (Linnaeus, 1768), Peripsocus phaeopterus (Stephens, 1836), Philotarsus picicornis (Fabricius, 1793), Amphigerontia bifasciata (Latreille, 1799), Psococerastis gibbosa (Sulzer, 1766), and Metylophorus nebulosus (Stephens, 1836). These species belong to five distantly related families of the largest psocid suborder Psocomorpha : Caeciliusidae , Stenopsocidae , Peripsocidae , Philotarsidae , and Psocidae . We show that all the examined species share a similar location of 18S rDNA on a medium-sized pair of autosomes. This is the first study of rDNA clusters in the order Psocoptera using FISH. We also demonstrate that these species have the classical insect (TTAGG) n telomere organization. Our results provide a foundation for further cytogenetic characterization and chromosome evolution studies in Psocoptera .
Related collections
Most cited references23
- Record: found
- Abstract: found
- Article: not found
Phylogenetic distribution of TTAGG telomeric repeats in insects.
- Record: found
- Abstract: found
- Article: not found
High dynamics of rDNA cluster location in kissing bug holocentric chromosomes (Triatominae, Heteroptera).
- Record: found
- Abstract: found
- Article: found