- Record: found
- Abstract: found
- Article: found
Structurally Conserved Primate LncRNAs Are Transiently Expressed during Human Cortical Differentiation and Influence Cell-Type-Specific Genes
Read this article at
Summary
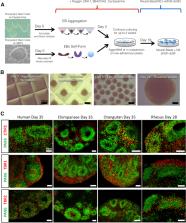
The cerebral cortex has expanded in size and complexity in primates, yet the molecular innovations that enabled primate-specific brain attributes remain obscure. We generated cerebral cortex organoids from human, chimpanzee, orangutan, and rhesus pluripotent stem cells and sequenced their transcriptomes at weekly time points for comparative analysis. We used transcript structure and expression conservation to discover gene regulatory long non-coding RNAs (lncRNAs). Of 2,975 human, multi-exonic lncRNAs, 2,472 were structurally conserved in at least one other species and 920 were conserved in all. Three hundred eighty-six human lncRNAs were transiently expressed (TrEx) and many were also TrEx in great apes (46%) and rhesus (31%). Many TrEx lncRNAs are expressed in specific cell types by single-cell RNA sequencing. Four TrEx lncRNAs selected based on cell-type specificity, gene structure, and expression pattern conservation were ectopically expressed in HEK293 cells by CRISPRa. All induced trans gene expression changes were consistent with neural gene regulatory activity.
Highlights
-
•
New orangutan and chimpanzee iPSC lines enable pan-primate gene expression analysis
-
•
Transiently expressed (TrEx) lncRNAs identified in primate cerebral cortex organoids
-
•
Conserved TrEx pattern correlates with cell-type specificity by single-cell RNA-seq
-
•
Four primate-conserved TrEx lncRNAs influence neural genes when ectopically expressed
Abstract
In this article, Salama and colleagues identified transiently expressed (TrEx) lncRNAs from human, chimpanzee, orangutan, and rhesus pluripotent stem cell-derived cerebral cortex organoids and assessed their structural and expression conservation. Transient expression correlated with cell-type specificity as measured by single-cell RNA-seq. Ectopic expression of TrEx lncRNAs by CRISPRa modulated expression of neural genes in trans, suggesting regulatory function.
Related collections
Most cited references24
- Record: found
- Abstract: found
- Article: not found
Organoids as an in vitro model of human development and disease.
- Record: found
- Abstract: found
- Article: not found
Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation.
- Record: found
- Abstract: found
- Article: not found