- Record: found
- Abstract: found
- Article: found
Inhibition of cyclooxygenase-1 by nonsteroidal anti-inflammatory drugs demethylates MeR2 enhancer and promotes Mbnl1 transcription in myogenic cells
Read this article at
Abstract
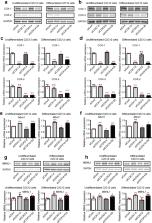
Muscleblind-like 1 (MBNL1) is a ubiquitously expressed RNA-binding protein, which is highly expressed in skeletal muscle. Abnormally expanded CUG-repeats in the DMPK gene cause myotonic dystrophy type 1 (DM1) by sequestration of MBNL1 to nuclear RNA foci and by upregulation of another RNA-binding protein, CUG-binding protein 1 (CUGBP1). We previously reported that a nonsteroidal anti-inflammatory drug (NSAID), phenylbutazone, upregulates MBNL1 expression in DM1 mouse model by demethylation of MeR2, an enhancer element in Mbnl1 intron 1. NSAIDs inhibit cyclooxygenase (COX), which is comprised of COX-1 and COX-2 isoforms. In this study, we screened 29 NSAIDs in C2C12 myoblasts, and found that 13 NSAIDs enhanced Mbnl1 expression, where COX-1-selective NSAIDs upregulated Mbnl1 more than COX-2-selective NSAIDs. Consistently, knockdown of COX-1, but not of COX-2, upregulated MBNL1 expression in C2C12 myoblasts and myotubes, as well as in myotubes differentiated from DM1 patient-derived induced pluripotent stem cells (iPSCs). Luciferase assay showed that COX-1-knockdown augmented the MeR2 enhancer activity. Furthermore, bisulfite sequencing analysis demonstrated that COX-1-knockdown suppressed methylation of MeR2. These results suggest that COX-1 inhibition upregulates Mbnl1 transcription through demethylation of the MeR2 enhancer. Taken together, our study provides new insights into the transcriptional regulation of Mbnl1 by the COX-1-mediated pathway.
Related collections
Most cited references55
- Record: found
- Abstract: found
- Article: not found
Transcription factors: from enhancer binding to developmental control.
- Record: found
- Abstract: found
- Article: not found