- Record: found
- Abstract: found
- Article: found
CRISPR-Cas Immunity against Phages: Its Effects on the Evolution and Survival of Bacterial Pathogens
research-article

Read this article at
There is no author summary for this article yet. Authors can add summaries to their articles on ScienceOpen to make them more accessible to a non-specialist audience.
Abstract
Introduction
Clustered regularly interspaced short palindromic repeats (CRISPR) loci are arrays
of short repeats separated by equally short “spacer” sequences [1]–[3]. Along with
the CRISPR-associated (cas) genes, they encode an adaptive immune system of archaea
and bacteria that protects the cell against viral infection [4]. Remarkably, this
system is capable of inserting a short piece of an infecting viral genome as a spacer
in the CRISPR array [4], [5] (Figure 1A). The spacer sequence is transcribed and processed
to generate a small antisense RNA (the CRISPR RNA or crRNA) (Figure 1B) [6] that is
used as a guide for the recognition and destruction of the invader in subsequent infections
(Figure 1C) [7]. Thus, spacer acquisition immunizes the bacterium and its progeny
against the virus from which it was taken. Because spacers are incorporated in sequential
order, CRISPR loci reflect the history of viral infection of the host. Cas proteins
participate in all the different steps of this pathway, namely the insertion of spacer
sequences into the CRISPR array [8], [9], the biogenesis of crRNAs [10], [11], and
the destruction of the infecting viral genome [12], [13].
10.1371/journal.ppat.1003765.g001
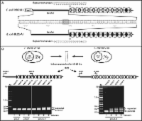
Figure 1
The CRISPR immunity pathway.
CRISPR loci contain clusters of repeats (white boxes) and spacers (colored boxes)
that are flanked CRISPR-associated (cas) genes. (A) During adaptation new spacers
derived from the genome of the invading virus are incorporated into the CRISPR array
by an unknown mechanism. Repeat duplication is also required. (B) During crRNA biogenesis
a CRISPR precursor transcript is processed by Cas endoribonucleases within repeat
sequences to generate small crRNAs. (C) During targeting the match between the crRNA
spacer and target sequences specifies the nucleolytic cleavage of invading mobile
genetic elements such as viruses and plasmids. (D) In the CRISPR-Cas system of F.
novicida, the tracrRNA (a small RNA mediated in crRNA biogenesis in this system) contains
homology to the BLP (bacterial lipoprotein) transcript. The base-pair interaction
between the tracrRNA and the BLP mRNA (mediated also by another small RNA, the scaRNA,
and the nuclease Cas9) regulates the expression of this immunomodulatory lipoprotein.
Distribution of CRISPR-Cas Loci among Bacterial Pathogens
In spite of the unique role that CRISPR-Cas loci play in antiviral defense, they are
not universal. To date, the CRISPR database [14], a webtool that determines the presence
of CRISPR arrays in completed genomes, indicates that 119/141 archaeal (84%) and 1012/2113
bacterial (48%) genomes contain CRISPR loci. In bacteria, there are species in which
all strains have CRISPR loci, some in which only some strains have these loci, and
species without strains having CRISPR loci. Therefore it is not possible to determine
unequivocally that lack of CRISPR in certain strains or species is due to loss of
these loci. However, because CRISPR sequences are spread thorough horizontal gene
transfer [15], [16] and can be easily lost [17]–[20], it has been hypothesized recently
that CRISPR are in a constant state of flux and can appear and disappear depending
on the selective forces of the environment [20]. The same type of uneven distribution
is found when we look at the presence of CRISPR loci in bacterial pathogens in the
CRISPR database (http://crispr.u-psud.fr/crispr/).
CRISPR-Cas Systems as a Barrier to Horizontal Gene Transfer
While most of the spacers with matches on GenBank target prokaryotic viruses (phages),
there is a still an important fraction that match other targets. A recent study looked
at all the spacer hits of archaeal CRISPR loci [21] and reported that 40% of them
matched phage sequences. The remaining 60% matched other mobile genetic elements such
as conjugative plasmids and transposons (22%), CRISPR-Cas loci (18%), and other genes
not associated with mobile elements (hypothetical ORFs and housekeeping genes, 20%).
Although an equally extensive study has not been performed with bacterial CRISPR spacers,
partial analysis suggests a similar distribution [22], [23]. While the presence of
antiphage spacers is key for the defense of the cell, the origin and function of these
nonphage targeting spacers is obscure. How are these spacers acquired? One possibility
is that these sequences are inserted into CRISPR loci during the transfer of foreign
genetic material that commonly occurs between prokaryotes, also known as horizontal
gene transfer (HGT) [24]. In this scenario, non-antiphage spacers are acquired during
bacteriophage transduction, plasmid conjugation, or upon the uptake of foreign DNA
during natural transformation. Alternatively, spacer acquisition only occurs as an
adaptive response to phage infection and the nonphage targeting spacers are acquired
only from phage transducing particles [25]. Regardless of whether the diversity of
the CRISPR spacer repertoire is generated by accident or not, the fact that CRISPR
loci can target all sorts of genetic material argues that these loci constitute a
barrier against the horizontal transfer of genes and accessory genetic elements. Indeed,
CRISPR interference has been shown experimentally to prevent the acquisition of conjugative
plasmids [26], integrative conjugative elements [27], and environmental DNA by natural
transformation [17], [28]. What is even more puzzling is the function, if any, of
these nonphage targeting spacers. Plasmid targeting could eliminate the burden of
additional replicating elements inside the cell, and the targeting of housekeeping
genes could provide a regulatory function for these spacers. However, plasmids, mobile
genetic elements, and foreign genes can provide a fitness advantage or even be essential
for survival (e.g., antibiotic resistance genes).
Implications of CRISPR-Mediated Targeting of Mobile Genetic Elements in Bacterial
Pathogens
HGT is the major source of genetic diversity for bacterial evolution [24]. In the
past century, the introduction of modern antibacterial therapies has accelerated the
evolution of pathogens. While it is clear that HGT has played a central role in the
spread of virulence factors and antimicrobial resistance genes [29], [30], only a
few studies have addressed whether and how CRISPR loci, owing to their potential to
regulate HGT, impact the evolution of pathogens. One of these studies investigated
the relationship between the CRISPR loci and the prophage content of group A streptococci
(GAS, Streptococcus pyogenes), one of the most prevalent human bacterial pathogens.
These organisms contain between two to eight prophages, each encoding at least one
virulence factor [31]. Bioinformatic analysis revealed that seven of the 13 available
GAS genomes contain CRISPR-Cas loci and that there is a mutually exclusive relationship
between CRISPR spacer sequences and their prophage targets [32]. This suggests that
there is a dynamic relationship between S. pyogenes, its phages, and its CRISPR loci
that results in the selection of strains with increased pathogenic adaptations. CRISPR-Cas
loci also can impact the spread of antibiotic resistance. Pathogenic staphylococci
have acquired resistance to all known antibiotics [33], primarily through the acquisition
of conjugative plasmids carrying resistance genes [30]. Staphylococcus epidermidis
RP62a is a clinical isolate containing a CRISPR-Cas system with a spacer matching
all staphylococcal conjugative plasmids sequenced to date [34]. This spacer provides
immunity against the conjugative transfer of these plasmids [26], thereby preventing
the acquisition of the antibiotic resistances that they carry. Therefore CRISPR loci
could control the dissemination of antibiotic resistance in staphylococci. This does
not seem to be the case for Escherichia coli. A study of a collection of 263 natural
E. coli isolates from human and animal hosts revealed that CRISPR loci neither match
plasmid sequences nor correlate with the presence or absence of plasmids or antibiotic
resistance genes [35].
Loss of CRISPR-Cas Loci in Bacterial Pathogens
CRISPR immunity against conjugative plasmids would compromise the survival of S. epidermidis
RP62a, and other staphylococci carrying similar CRISPR-Cas systems [36], [37] in hospital
or other settings where antibiotics are used. A recent study [20] looked for the transfer
of the mupirocin-resistant conjugative plasmid pG0400 into S. epidermidis to determine
if a CRISPR-Cas system and its target could coexist to prevent this potentially detrimental
antiplasmid activity of CRISPR immunity. Immunity against the plasmid was found to
decrease the transfer efficiency by about four orders of magnitude but not absolute.
Transconjugants that evaded CRISPR attack were analyzed only to find that in all cases
they harbored preexisting CRISPR-Cas mutations that allowed plasmid transfer. Loss
of CRISPR-Cas loci upon transfer of antibiotic resistant plasmids also seems to occur
in enterococci. A screen of 45 strains of Enterococcus faecalis showed a correlation
between the presence of CRISPR-Cas loci and antibiotic resistance genes [38]. Finally,
another recent study explored the consequences of CRISPR targeting of Streptococcus
pneumoniae capsule genes, essential for pneumococcal infection. During infection,
natural transformation of capsule genes allows nonencapsulated, avirulent pneumococci
to become encapsulated and kill the mice [39]. A CRISPR-Cas targeting a specific capsule
gene was engineered into nonencapsulated S. pneumoniae and used to infect mice in
the presence of heat-killed encapsulated pneumococci [17]. Horizontal transfer of
capsule genes from heat-killed cells into live, nonencapsulated bacteria was prevented
by CRISPR immunity, resulting in the survival of mice. The occasional mice that succumbed
to pneumococcal infection, however, contained encapsulated bacteria carrying inactivating
mutations in the engineered CRISPR locus. These and other results [18], [19] suggest
that CRISPR loci and their targets cannot coexist in the same cell. In the case of
strong environmental selection of a targeted gene or mobile element, only CRISPR mutants
survive. This is a possible explanation for the lack of CRISPR in S. pneumoniae and
S. aureus, two notoriously fast-evolving pathogens, but also in other bacteria and
archaea that lack this immune system.
A Direct Role for CRISPR-Cas Systems in Bacterial Pathogenesis
While the reasons for the absence of CRISPR-Cas loci in some fast-evolving pathogens
remain a matter of speculation, recent evidence showed that these loci can also promote
pathogenesis. A study in Legionella pneumophila showed that cas2, a gene involved
in the acquisition of new spacers, is required for the propagation of this pathogen
inside amoebae hosts [40], although it is not clear what the function of this gene
is during growth. More compelling evidence is found in the intracellular pathogen
Francisella novicida. In this bacterium, cas9 is a CRISPR-associated dsDNA nuclease
that requires, in addition to the crRNA guide, a tracrRNA (trans-activating crRNA)
for cleavage of the invader genome [41], [42]. It was found recently that cas9 is
required to repress the production of a bacterial lipoprotein (BLP), a toll-like receptor
2 (TLR2) ligand that induces an innate immune inflammatory response [43]. Repression
is independent of the crRNA guides, but requires the tracrRNA and a new small CRISPR-associated
RNA (scaRNA) with complementarity to the tracrRNA [44], [45]. The tracrRNA, in turn,
contains an ∼85 nt region with partial complementarity to the 3′-end of the BLP messenger,
an interaction that leads to the BLP mRNA degradation through an unknown mechanism.
This CRISPR-mediated regulation of BLP expression allows F. novicida to evade the
host's immune response. A similar mechanism seems to be in place in other pathogens
as well: deletion of cas9 in Neisseria meningitidis affected virulence traits such
as adherence to and invasion of human epithelial cells [44], and inactivation of cas9
in Campylobacter jejuni resulted in reduced virulence [46]. While the predominance
of tracrRNA/scaRNA-mediated regulation remains to be investigated, its existence suggests
that CRISPR-Cas loci can be easily converted into regulatory elements that enhance
bacterial pathogenesis.
Conclusions
Clearly CRISPR-Cas systems can both prevent the evolution of pathogenesis, and thus
be lost or mutated in bacterial pathogens, but also be co-opted by the pathogen to
increase virulence. This will depend of a series of factors: whether other antiphage
systems can fulfill the function of the lost CRISPR-Cas system, whether the pathogen
relies heavily on HGT for survival, and whether the CRISPR-Cas system can be easily
converted into a regulator of gene expression. In the face of the lateral transfer
of CRISPR systems, the repression of gene expression by CRISPR provides another level
of selection for the maintenance of these systems. While the repression of BLP provides
a selectable advantage for Francisella, the accidental repression of essential genes
(which could be produced by a fortuitous base-pairing of the tracrRNA and an essential
transcript) will select against the lateral transfer of some CRISPR-Cas systems into
certain hosts. In the future, the application of DNA sequencing technologies to epidemiological
studies will allow us to measure correlations between the flux of CRISPR-Cas loci
and the acquisition of antibiotic-resistance plasmids and pathogenicity islands or
genes, thus allowing us to measure the effect of CRISPR on the emergence of virulence.
On the other hand, the importance of CRISPR for pathogenesis provides a new target
for antimicrobials with anti-CRISPR activity. Interestingly, phages already found
such anti-CRISPR compounds for us: as part of their arms race with bacteria, phages
have developed CRISPR inhibitors [47]. The intersection between CRISPR biology and
bacterial pathogenesis is a new and exciting research area that is only beginning
to be explored.
Related collections
Most cited references32
- Record: found
- Abstract: found
- Article: not found
CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA.
Luciano A. Marraffini, Erik Sontheimer (2008)

- Record: found
- Abstract: found
- Article: found
The CRISPRdb database and tools to display CRISPRs and to generate dictionaries of spacers and repeats
- Record: found
- Abstract: found
- Article: found
Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli
Ido Yosef, Moran Goren, Udi Qimron (2012)