- Record: found
- Abstract: found
- Article: found
Cellular taxonomy and spatial organization of the murine ventral posterior hypothalamus

Read this article at
Abstract
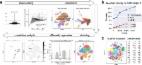
The ventral posterior hypothalamus (VPH) is an anatomically complex brain region implicated in arousal, reproduction, energy balance, and memory processing. However, neuronal cell type diversity within the VPH is poorly understood, an impediment to deconstructing the roles of distinct VPH circuits in physiology and behavior. To address this question, we employed a droplet-based single-cell RNA sequencing (scRNA-seq) approach to systematically classify molecularly distinct cell populations in the mouse VPH. Analysis of >16,000 single cells revealed 20 neuronal and 18 non-neuronal cell populations, defined by suites of discriminatory markers. We validated differentially expressed genes in selected neuronal populations through fluorescence in situ hybridization (FISH). Focusing on the mammillary bodies (MB), we discovered transcriptionally-distinct clusters that exhibit neuroanatomical parcellation within MB subdivisions and topographic projections to the thalamus. This single-cell transcriptomic atlas of VPH cell types provides a resource for interrogating the circuit-level mechanisms underlying the diverse functions of VPH circuits.
Related collections
Most cited references118
- Record: found
- Abstract: found
- Article: not found
Fast, sensitive, and accurate integration of single cell data with Harmony
- Record: found
- Abstract: found
- Article: found
SCANPY : large-scale single-cell gene expression data analysis

- Record: found
- Abstract: found
- Article: found