- Record: found
- Abstract: found
- Article: found
In BPS1 Downregulated Roots, the BYPASS1 Signal Disrupts the Induction of Cortical Cell Divisions in Bean- Rhizobium Symbiosis
Read this article at
Abstract
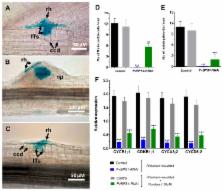
BYPASS1 ( BPS1), which is a well-conserved gene in plants, is required for normal root and shoot development. In the absence of BPS1 gene function, Arabidopsis overproduces a mobile signalling compound (the BPS1 signal) in roots, and this transmissible signal arrests shoot growth and causes abnormal root development. In addition to the shoot and root meristem activities, the legumes also possess transient meristematic activity in root cortical cells during Rhizobium symbiosis. We explored the role of Phaseolus vulgaris BPS1 during nodule primordium development using an RNA-interference (RNAi) silencing approach. Our results show that upon Rhizobium infection, the PvBPS1-RNAi transgenic roots failed to induce cortical cell divisions without affecting the rhizobia-induced root hair curling and infection thread formation. The transcript accumulation of early nodulin genes, cell cyclins, and cyclin-dependent kinase genes was affected in RNAi lines. Interestingly, the PvBPS1-RNAi root nodule phenotype was partially rescued by exogenous application of fluridone, a carotenoid biosynthesis inhibitor, which was used because the carotenoids are precursors of BPS1 signalling molecules. Furthermore, we show that the PvBPS1 promoter was active in the nodule primordia. Together, our data show that PvBPS1 plays a vital role in the induction of meristematic activity in root cortical cells and in the establishment of nodule primordia during Phaseolus-Rhizobium symbiosis.
Related collections
Most cited references48
- Record: found
- Abstract: found
- Article: not found
Strigolactone inhibition of shoot branching.
- Record: found
- Abstract: not found
- Article: not found
Assaying chimeric genes in plants: The GUS gene fusion system
- Record: found
- Abstract: found
- Article: not found