- Record: found
- Abstract: found
- Article: found
A Probabilistic Matrix Factorization Method for Identifying lncRNA-Disease Associations
Read this article at
Abstract
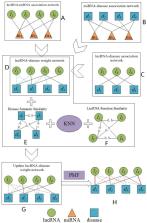
Recently, an increasing number of studies have indicated that long-non-coding RNAs (lncRNAs) can participate in various crucial biological processes and can also be used as the most promising biomarkers for the treatment of certain diseases such as coronary artery disease and various cancers. Due to costs and time complexity, the number of possible disease-related lncRNAs that can be verified by traditional biological experiments is very limited. Therefore, in recent years, it has been very popular to use computational models to predict potential disease-lncRNA associations. In this study, we constructed three kinds of association networks, namely the lncRNA-miRNA association network, the miRNA-disease association network, and the lncRNA-disease correlation network firstly. Then, through integrating these three newly constructed association networks, we constructed an lncRNA-disease weighted association network, which would be further updated by adopting the KNN algorithm based on the semantic similarity of diseases and the similarity of lncRNA functions. Thereafter, according to the updated lncRNA-disease weighted association network, a novel computational model called PMFILDA was proposed to infer potential lncRNA-disease associations based on the probability matrix decomposition. Finally, to evaluate the superiority of the new prediction model PMFILDA, we performed Leave One Out Cross-Validation (LOOCV) based on strongly validated data filtered from MNDR and the simulation results indicated that the performance of PMFILDA was better than some state-of-the-art methods. Moreover, case studies of breast cancer, lung cancer, and colorectal cancer were implemented to further estimate the performance of PMFILDA, and simulation results illustrated that PMFILDA could achieve satisfying prediction performance as well.
Related collections
Most cited references37

- Record: found
- Abstract: found
- Article: found
Long non-coding RNAs and complex diseases: from experimental results to computational models
- Record: found
- Abstract: found
- Article: not found
DD3: a new prostate-specific gene, highly overexpressed in prostate cancer.

- Record: found
- Abstract: found
- Article: found