- Record: found
- Abstract: found
- Article: found
Mapping of ESE-1 subdomains required to initiate mammary epithelial cell transformation via a cytoplasmic mechanism
Read this article at
Abstract
Background
The ETS family transcription factor ESE-1 is often overexpressed in human breast cancer. ESE-1 initiates transformation of MCF-12A cells via a non-transcriptional, cytoplasmic process that is mediated by a unique 40-amino acid serine and aspartic acid rich (SAR) subdomain, whereas, ESE-1's nuclear transcriptional property is required to maintain the transformed phenotype of MCF7, ZR-75-1 and T47D breast cancer cells.
Results
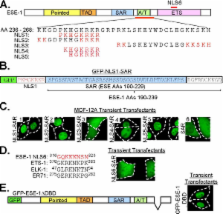
To map the minimal functional nuclear localization (NLS) and nuclear export (NES) signals, we fused in-frame putative NLS and NES motifs between GFP and the SAR domain. Using these GFP constructs as reporters of subcellular localization, we mapped a single NLS to six basic amino acids ( 242HGKRRR 247) in the AT-hook and two CRM1-dependent NES motifs, one to the pointed domain (NES1: 102LCNCALEELRL 112) and another to the DNA binding domain (DBD), (NES2: 275LWEFIRDILI 284). Moreover, analysis of a putative NLS located in the DBD ( 316GQKKKNSN 323) by a similar GFP-SAR reporter or by internal deletion of the DBD, revealed this sequence to lack NLS activity. To assess the role of NES2 in regulating ESE-1 subcellular localization and subsequent transformation potency, we site-specifically mutagenized NES2, within full-length GFP-ESE-1 and GFP-NES2-SAR reporter constructs. These studies show that site-specific mutation of NES2 completely abrogates ESE-1 transforming activity. Furthermore, we show that exclusive cytoplasmic targeting of the SAR domain is sufficient to initiate transformation, and we report that an intact SAR domain is required, since block mutagenesis reveals that an intact SAR domain is necessary to maintain its full transforming potency. Finally, using a monoclonal antibody targeting the SAR domain, we demonstrate that the SAR domain contains a region accessible for protein - protein interactions.
Conclusions
These data highlight that ESE-1 contains NLS and NES signals that play a critical role in regulating its subcellular localization and function, and that an intact SAR domain mediates MEC transformation exclusively in the cytoplasm, via a novel nontranscriptional mechanism, whereby the SAR motif is accessible for ligand and/or protein interactions. These findings are significant, since they provide novel molecular insights into the functions of ETS transcription factors in mammary cell transformation.
Related collections
Most cited references36
- Record: found
- Abstract: found
- Article: not found
The ETS-domain transcription factor family.
- Record: found
- Abstract: found
- Article: not found
Molecular biology of the Ets family of transcription factors.
- Record: found
- Abstract: found
- Article: not found