- Record: found
- Abstract: found
- Article: found
Integrated analysis of numerous heterogeneous gene expression profiles for detecting robust disease-specific biomarkers and proposing drug targets

Read this article at
Abstract
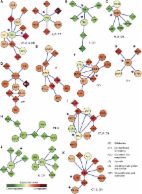
Genome-wide expression profiling has revolutionized biomedical research; vast amounts of expression data from numerous studies of many diseases are now available. Making the best use of this resource in order to better understand disease processes and treatment remains an open challenge. In particular, disease biomarkers detected in case–control studies suffer from low reliability and are only weakly reproducible. Here, we present a systematic integrative analysis methodology to overcome these shortcomings. We assembled and manually curated more than 14 000 expression profiles spanning 48 diseases and 18 expression platforms. We show that when studying a particular disease, judicious utilization of profiles from other diseases and information on disease hierarchy improves classification quality, avoids overoptimistic evaluation of that quality, and enhances disease-specific biomarker discovery. This approach yielded specific biomarkers for 24 of the analyzed diseases. We demonstrate how to combine these biomarkers with large-scale interaction, mutation and drug target data, forming a highly valuable disease summary that suggests novel directions in disease understanding and drug repurposing. Our analysis also estimates the number of samples required to reach a desired level of biomarker stability. This methodology can greatly improve the exploitation of the mountain of expression profiles for better disease analysis.
Related collections
Most cited references49
- Record: found
- Abstract: found
- Article: found
Network-based classification of breast cancer metastasis

- Record: found
- Abstract: found
- Article: found
IntAct—open source resource for molecular interaction data

- Record: found
- Abstract: found
- Article: found