- Record: found
- Abstract: found
- Article: found
Targeting circular RNAs as a therapeutic approach: current strategies and challenges
Read this article at
Abstract
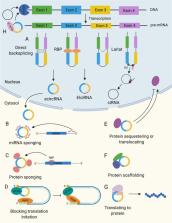
Significant progress has been made in circular RNA (circRNA) research in recent years. Increasing evidence suggests that circRNAs play important roles in many cellular processes, and their dysregulation is implicated in the pathogenesis of various diseases. CircRNAs are highly stable and usually expressed in a tissue- or cell type-specific manner. Therefore, they are currently being explored as potential therapeutic targets. Gain-of-function and loss-of-function approaches are typically performed using circRNA expression plasmids and RNA interference-based strategies, respectively. These strategies have limitations that can be mitigated using nanoparticle and exosome delivery systems. Furthermore, recent developments show that the cre-lox system can be used to knockdown circRNAs in a cell-specific manner. While still in the early stages of development, the CRISPR/Cas13 system has shown promise in knocking down circRNAs with high specificity and efficiency. In this review, we describe circRNA properties and functions and highlight their significance in disease. We summarize strategies that can be used to overexpress or knockdown circRNAs as a therapeutic approach. Lastly, we discuss major challenges and propose future directions for the development of circRNA-based therapeutics.
Related collections
Most cited references165
- Record: found
- Abstract: found
- Article: not found
Genome engineering using the CRISPR-Cas9 system.

- Record: found
- Abstract: found
- Article: found
Double-slit photoelectron interference in strong-field ionization of the neon dimer
- Record: found
- Abstract: found
- Article: not found