- Record: found
- Abstract: found
- Article: found
Entrainment to Periodic Initiation and Transition Rates in a Computational Model for Gene Translation
Read this article at
Abstract
Periodic oscillations play an important role in many biomedical systems. Proper functioning
of biological systems that respond to periodic signals requires the ability to synchronize
with the periodic excitation. For example, the sleep/wake cycle is a manifestation
of an internal timing system that synchronizes to the solar day. In the terminology
of systems theory, the biological system must entrain or phase-lock to the periodic
excitation. Entrainment is also important in synthetic biology. For example, connecting
several artificial biological systems that entrain to a common clock may lead to a
well-functioning modular system. The cell-cycle is a periodic program that regulates
DNA synthesis and cell division. Recent biological studies suggest that cell-cycle
related genes entrain to this periodic program at the gene translation level, leading
to periodically-varying protein levels of these genes. The ribosome flow model (RFM)
is a deterministic model obtained via a mean-field approximation of a stochastic model
from statistical physics that has been used to model numerous processes including
ribosome flow along the mRNA. Here we analyze the RFM under the assumption that the
initiation and/or transition rates vary periodically with a common period
 . We show that the ribosome distribution profile in the RFM entrains to this periodic
excitation. In particular, the protein synthesis pattern converges to a unique periodic
solution with period
. We show that the ribosome distribution profile in the RFM entrains to this periodic
excitation. In particular, the protein synthesis pattern converges to a unique periodic
solution with period
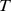 . To the best of our knowledge, this is the first proof of entrainment in a mathematical
model for translation that encapsulates aspects such as initiation and termination
rates, ribosomal movement and interactions, and non-homogeneous elongation speeds
along the mRNA. Our results support the conjecture that periodic oscillations in tRNA
levels and other factors related to the translation process can induce periodic oscillations
in protein levels, and may suggest a new approach for re-engineering genetic systems
to obtain a desired, periodic, protein synthesis rate.
. To the best of our knowledge, this is the first proof of entrainment in a mathematical
model for translation that encapsulates aspects such as initiation and termination
rates, ribosomal movement and interactions, and non-homogeneous elongation speeds
along the mRNA. Our results support the conjecture that periodic oscillations in tRNA
levels and other factors related to the translation process can induce periodic oscillations
in protein levels, and may suggest a new approach for re-engineering genetic systems
to obtain a desired, periodic, protein synthesis rate.
Related collections
Most cited references58
- Record: found
- Abstract: found
- Article: not found
Mistranslation-induced protein misfolding as a dominant constraint on coding-sequence evolution.
- Record: found
- Abstract: found
- Article: not found
Synthetic biology: applications come of age
- Record: found
- Abstract: found
- Article: not found