- Record: found
- Abstract: found
- Article: found
Plumbagin inhibits proliferation and induces apoptosis of hepatocellular carcinoma by downregulating the expression of SIVA
Abstract
Purpose: Plumbagin is thought to be a bioactive phytochemical drug and exerts an antitumor effect on various cancers. However, few studies focus on the antitumor activity of plumbagin on liver cancer. This study first investigated the antitumor activity of plumbagin on liver cancer and further investigated the molecular mechanism of its antitumor activity against hepatocellular carcinoma, both in vitro and in vivo.
Methods: The antiproliferative activity of plumbagin was evaluated through CCK-8, EdU, and colony forming test. The cell cycle and apoptosis were then analyzed by flow cytometer. Western blot was used to detect the expression of apoptosis related protein, SIVA, and mTOR pathway. RNA-seq was performed to determine the gene expression profiles and overexpressed or knocked down SIVA to validate its role in plumbagin’s antitumor activity. Regarding animal experiment, a xenograft model in BALB/c nude mice was built using LM3-Luci cells. Then bioluminescence imaging and further immunohistochemistry were performed to study the antitumor activity and the expression of SIVA and mTOR in the plumbagin-treated group.
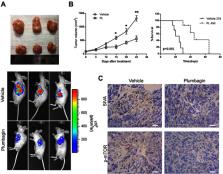
Results: Plumbagin can inhibit proliferation and induce apoptosis of liver cancer cells in vitro. Further experiment demonstrated that plumbagin could inhibit the expression of SIVA and subsequently downregulate the mTOR signaling pathway, and upregulating the expression of SIVA will alleviate the antitumor activity of plumbagin on liver cancer, which confirmed the important role of the SIVA/mTOR signaling pathway in the antitumor activity of plumbagin. In vivo bioluminescence imaging showed a decreased signal in the plumbagin-treated group, and further immunohistochemistry demonstrated that plumbagin could inhibit the SIVA/mTOR signaling pathway in tumor tissues.
Conclusion: Our promising results showed that plumbagin could inhibit proliferation and induce apoptosis of hepatic cancer through inhibiting the SIVA/mTOR signaling pathway for the first time, which indicated that plumbagin might be a good candidate against liver cancer.
Most cited references27
- Record: found
- Abstract: found
- Article: not found
RAFT1: a mammalian protein that binds to FKBP12 in a rapamycin-dependent fashion and is homologous to yeast TORs.

- Record: found
- Abstract: found
- Article: found
The mTOR Signalling Pathway in Human Cancer
- Record: found
- Abstract: found
- Article: not found
