- Record: found
- Abstract: found
- Article: found
Diversity and relative abundance of the bacterial pathogen, Flavobacterium spp., infecting reproductive ecotypes of kokanee salmon
Read this article at
Abstract
Background
Understanding the distribution and abundance of pathogens can provide insight into the evolution and ecology of their host species. Previous research in kokanee, the freshwater form of sockeye salmon ( Oncorhynchus nerka), found evidence that populations spawning in streams may experience a greater pathogen load compared with populations that spawn on beaches. In this study we tested for differences in the abundance and diversity of the gram-negative bacteria, Flavobacterium spp., infecting tissues of kokanee in both of these spawning habitats (streams and beaches). Molecular assays were carried out using primers designed to amplify a ~200 nucleotide region of the gene encoding the ATP synthase alpha subunit ( AtpA) within the genus Flavobacterium. Using a combination of DNA sequencing and quantitative PCR (qPCR) we compared the diversity and relative abundance of Flavobacterium AtpA amplicons present in DNA extracted from tissue samples of kokanee collected from each spawning habitat.
Results
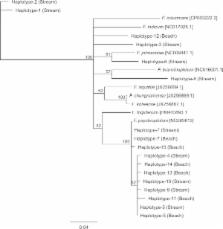
We identified 10 Flavobacterium AtpA haplotypes among the tissues of stream-spawning kokanee and seven haplotypes among the tissues of beach-spawning kokanee, with only two haplotypes shared between spawning habitats. Haplotypes occurring in the same clade as F. psychrophilum were the most prevalent (92% of all reads, 60% of all haplotypes), and occurred in kokanee from both spawning habitats (streams and beaches). Subsequent qPCR assays did not find any significant difference in the relative abundance of Flavobacterium AtpA amplicons between samples from the different spawning habitats.
Conclusions
We confirmed the presence of Flavobacterium spp. in both spawning habitats and found weak evidence for increased Flavobacterium diversity in kokanee sampled from stream-spawning sites. However, the quantity of Flavobacterium DNA did not differ between spawning habitats. We recommend further study aimed at quantifying pathogen diversity and abundance in population-level samples of kokanee combined with environmental sampling to better understand the ecology of pathogen infection in this species.
Related collections
Most cited references33
- Record: found
- Abstract: found
- Article: not found
Cross-biome metagenomic analyses of soil microbial communities and their functional attributes.
- Record: found
- Abstract: not found
- Article: not found
R: A Language and environmental for statistical computing
- Record: found
- Abstract: not found
- Article: not found