- Record: found
- Abstract: found
- Article: found
BISMA - Fast and accurate bisulfite sequencing data analysis of individual clones from unique and repetitive sequences
Read this article at
Abstract
Background
Bisulfite sequencing is a popular method to analyze DNA methylation patterns at high resolution. A region of interest is targeted by PCR and about 20-50 subcloned DNA molecules are usually analyzed, to determine the methylation status at single CpG sites and molecule resolution.
Results
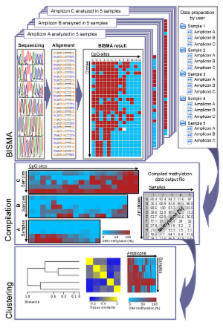
The BISMA (Bisulfite Sequencing DNA Methylation Analysis) software for analysis of primary bisulfite sequencing data implements sequencing data extraction and enhanced data processing, quality controls, analysis and presentation of the methylation state. It uses an improved strategy for detection of clonal molecules and accurate CpG site detection and it supports for the first time analysis of repetitive sequences.
Conclusions
BISMA works highly automated but still provides the user full control over all steps of the analysis. The BISMA software is freely available as an online tool for academic purposes for the analysis of bisulfite sequencing data from both unique and repetitive sequences http://biochem.jacobs-university.de/BDPC/BISMA/.
Related collections
Most cited references16
- Record: found
- Abstract: found
- Article: not found
A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands.
- Record: found
- Abstract: found
- Article: not found
A simple method for estimating global DNA methylation using bisulfite PCR of repetitive DNA elements.
- Record: found
- Abstract: found
- Article: not found