- Record: found
- Abstract: found
- Article: found
Footprints of genetic susceptibility to pulmonary tuberculosis: Cytokine gene variants in north Indians
Read this article at
Abstract
Background & objectives:
Tuberculosis is (TB) responsible for high morbidity and mortality worldwide. Cytokines play a major role in defense against Mycobacterium tuberculosis infection. Polymorphisms in the genes encoding the various pro- and anti-inflammatory cytokines have been associated with tuberculosis susceptibility. In this study we examined association of 25 sequence polymorphisms in six candidate cytokine genes namely IFNG, TNFB, IL4, IL1RA, IL1B and IL12 and their related haplotypes with risk of developing pulmonary tuberculosis (PTB) among north Indians.
Methods:
Pulmonary TB (n=110) patients and 215 healthy controls (HC) from north India were genotyped. Purified multiplex PCR products were subjected to mass spectrometry using Sequenom MassARRAY platform to generate the genotypes in a population-based case-control study.
Results:
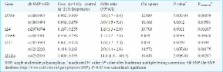
Using multiple corrections, significant overall risk against PTB was observed at seven loci which included variants in IFNG at rs1861493 and rs1861494; IL1RA at rs4252019, IL4 variant rs2070874, IL12 variants rs3212220, rs2853694 and TNFB variant rs1041981. Analysis of gene structure revealed two haplotype blocks formed by IFNG variants rs1861493 and rs1861494. The TA haplotype was significantly over-represented ( P=0.011) in the cases showing a two-fold risk in the current population (Odds ratio=1.59 CI=1.101 to 2.297) and TNFB variants at rs2229094 and rs1041981 contributed to two haplotypes which were in strong linkage disequilibrium (LD) with AT haplotype showing a three-fold risk ( P=0.0011, Odds ratio=3, CI=0.1939 to 0.7445) of developing PTB in north Indians.
Related collections
Most cited references26
- Record: found
- Abstract: found
- Article: not found
A mutation in the interferon-gamma-receptor gene and susceptibility to mycobacterial infection.
- Record: found
- Abstract: found
- Article: not found
Global epidemiology of tuberculosis. Morbidity and mortality of a worldwide epidemic.
- Record: found
- Abstract: found
- Article: not found