- Record: found
- Abstract: found
- Article: found
Editorial: Antimicrobial resistance and one health: from culture to genomics
editorial
Read this article at
There is no author summary for this article yet. Authors can add summaries to their articles on ScienceOpen to make them more accessible to a non-specialist audience.
Abstract
Antimicrobial resistance (AMR) is among the leading global public health threats of
the 21st century. Thus, AMR threatens human, animal, and environmental health globally
(Abia and Essack, 2023). Moreover, the widespread misuse and uncontrolled prescription
of antibiotics aggravates the problem, diminishing antibiotic efficacy, including
last-resort ones (Lu et al., 2023). This problem is accentuated in rural settings
in low- and middle-income countries where community members are small-scale farmers,
often living with their livestock inside the yards and lacking adequate water and
sanitation facilities (Sulis et al., 2022). Inadequate animal and human waste disposal
from household and agricultural practices could transfer antimicrobial-resistant pathogens
and their associated genes to the environment. These pollutants may pose a health
risk to resource-limited community members upon exposure to such polluted environments.
Therefore, monitoring antimicrobial resistance requires a one health approach involving
the human, animal, and environmental sectors. Furthermore, employing advanced genomic
tools to complement existing culture-based techniques would provide a more comprehensive
appreciation of the extent of AMR globally. Therefore, the current Research Topic
aimed at providing an update on the one health approach to AMR using cultural and
genomic approaches. This Research Topic includes seven articles by 45 authors from
eight countries in the Americas, Asia, and Europe.
The development of resistance in bacteria is a complex process involving several mechanisms.
Ramamurthy et al. reviewed the genetic network and programmed regulations used by
bacterial pathogens in antimicrobial resistance. Mechanisms identified could be cellular
(outer membrane modifications, enzyme inactivation, biofilm formation, and efflux
pump activation) and/or molecular (gene mutations and upregulation, quorum-sensing
networks, and other mechanisms). Once developed, mobile genetic elements (e.g., plasmids,
integrons, and insertion sequences) mostly aid in disseminating AMR in the environment,
influenced by the presence of stressors such as biocides, antibiotics, heavy metals,
and detergents.
Children are vulnerable to infections due to their less developed immune systems.
Patil et al. assessed the resistance genomics and molecular epidemiology of Pseudomonas
aeruginosa in young children, specifically targeting high-risk extended-spectrum beta-lactamase
(ESBL)-producing clones. The authors analysed 294 clinical isolates from a paediatric
hospital in China using culture-dependent and independent methods and found that 56%,
40%, 39%, 36%, 33%, and 32% of the isolates were resistant to piperacillin-tazobactam,
cefepime, ceftazidime, imipenem, meropenem, and ciprofloxacin, respectively. Furthermore,
42% of the isolates were ESBL producers. Polymerase chain reaction and sequencing
revealed the presence of numerous genes including bla
CTX-M-15, bla
NDM-1, aac(3)IIIa, tet(A), and 23 sequence types, as well as a de novo strain. The
isolates also harboured various plasmids that could aid in the transmission of these
resistance genes. Still in humans, Jia et al. genomically characterised bacteraemia-associated
carbapenem-resistant Serratia marcescens isolates and reported the emergence of KPC-2-encoding
IncR plasmids in a clinical setting to understand their resistance and transmission
dynamics. After sequencing the complete genome of two S. marcescens isolates, the
authors observed that the isolates harboured bla
KPC-2-bearing IncR plasmids and multiple plasmid-borne antimicrobial resistance genes.
Comparative plasmid analysis further suggested common ancestral lineage for both isolates
and concluded that the plasmid borne by both isolates could hinder the transmission
of KPC-2-producing S. marcescens in clinical settings.
While some clinical bacteria may acquire resistance traits due to exposure to external
stressors, others are intrinsically resistant to several antimicrobials. Hence, Shi
et al. functionally characterised a novel aminoglycoside phosphotransferase, APH(9)-Ic,
and its variant from Stenotrophomonas maltophilia. After performing recombinant protein
expression and enzyme kinetic studies, the authors used whole-genome sequencing to
determine the genetic context of the studied gene. The authors observed that aph(9)-Ic
and aph(9)-Ic1 conferred spectinomycin resistance. Furthermore, a recombinant strain
harbouring aph(9)-Ic displayed a marked increased minimum inhibitory concentration
level against spectinomycin compared with the control strains; aph(9)-Ic was chromosome-borne,
with a relatively conserved genetic environment and no mobile genetic element in its
surrounding region.
The one health approach has been shown to provide a comprehensive understanding of
the spread of AMR in the human–animal–environment triad. Thus, Yasmeen et al. investigated
the occurrence of ESBL-harbouring Klebsiella pneumoniae from 793 samples collected
from animals, humans, and the environment. Their study found K. pneumoniae in 11.6%,
8.4% and 7.0%, of animal, humans, and environmental samples, respectively, with the
animal isolates harbouring more ESBL genes (mostly bla
SHV) than their human and environmental counterparts. These results show the potential
spread of these pathogens within the three one health compartments.
Given the debating effects of AMR, involving humans, animals, and the environment,
attempts at curbing this ill must take a holistic approach involving multisectoral
stakeholders. Thus, Ahmad et al. reviewed how the one health concept has been implanted
and evolved in combating AMR. They identified that pillars of support to combat AMR
include instituting robust ABR surveillance in the various sectors individually and
in combination and at national and international level, as well as improving laboratory
resources and executing and strictly implementing core plans and actions. Furthermore,
drawing on lessons from the recent COVID-19 pandemic, Mantegazza et al. proposed a
de novo approach based on a convergence model to fight AMR, exploiting the current
Sustainable Development Goals (SDG) roadmap, emphasising a circular approach and highlighting
win–win scenarios by enforcing existing objectives and targets to strengthen a multidisciplinary
synergistic effort towards fighting AMR. They argued that the global importance of
AMR warrants that it be an integral part of the SDG roadmap, involving all the SDG
objectives, as such efforts could significantly advance health as a system by focusing
on a major priority such as AMR.
Collectively, these studies contribute to knowledge on the spread of AMR in humans,
animals, and the environment. Furthermore, they reveal the various mechanisms involved
in AMR in bacteria, including factors triggering their occurrence and factors favouring
such spread. Therefore, combining culture and advanced molecular techniques such as
genomics would paint a brighter picture regarding the overall AMR burden. Put together,
the studies provide evidence that adopting a one health concept for a successful fight
against AMR is imperative.
Author contributions
AA: Conceptualization, Methodology, Writing – original draft, Writing – review & editing.
AT: Conceptualization, Methodology, Writing – review & editing. NP: Conceptualization,
Methodology, Writing – review & editing.
Related collections
Most cited references3
- Record: found
- Abstract: found
- Article: not found
Antimicrobial resistance in low- and middle-income countries: current status and future directions
Giorgia Sulis, Sena Sayood, Sumanth Gandra (2021)
- Record: found
- Abstract: found
- Article: found
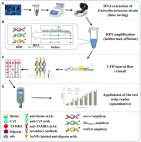
Rapid detection of multiple resistance genes to last-resort antibiotics in Enterobacteriaceae pathogens by recombinase polymerase amplification combined with lateral flow dipstick
Chenze Lu, Jingwen Wang, Leiming Pan … (2023)
Author and article information
Comments
Comment on this article
scite_
Similar content1,740
- EditorialAuthors:
- Editorial boardAuthors:
- Editorial working groupAuthors: