- Record: found
- Abstract: found
- Article: found
Development of a multi-locus CRISPR gene drive system in budding yeast
Read this article at
Abstract
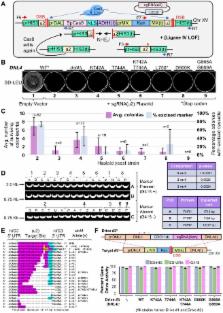
The discovery of CRISPR/Cas gene editing has allowed for major advances in many biomedical disciplines and basic research. One arrangement of this biotechnology, a nuclease-based gene drive, can rapidly deliver a genetic element through a given population and studies in fungi and metazoans have demonstrated the success of such a system. This methodology has the potential to control biological populations and contribute to eradication of insect-borne diseases, agricultural pests, and invasive species. However, there remain challenges in the design, optimization, and implementation of gene drives including concerns regarding biosafety, containment, and control/inhibition. Given the numerous gene drive arrangements possible, there is a growing need for more advanced designs. In this study, we use budding yeast to develop an artificial multi-locus gene drive system. Our minimal setup requires only a single copy of S. pyogenes Cas9 and three guide RNAs to propagate three gene drives. We demonstrate how this system could be used for targeted allele replacement of native genes and to suppress NHEJ repair systems by modifying DNA Ligase IV. A multi-locus gene drive configuration provides an expanded suite of options for complex attributes including pathway redundancy, combatting evolved resistance, and safeguards for control, inhibition, or reversal of drive action.
Related collections
Most cited references36

- Record: found
- Abstract: found
- Article: found
BLAST-EXPLORER helps you building datasets for phylogenetic analysis
- Record: found
- Abstract: found
- Article: not found
Repair of double-strand breaks by end joining.
- Record: found
- Abstract: found
- Article: not found
