scite_
- Record: found
- Abstract: found
- Poster: found
Reduced models for uncertainty quantification in the cardiovascular network via domain decomposition
poster
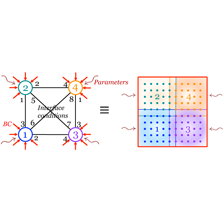
Abstract
Reduced 1D models of the cardiovascular system are widely employed to study the propagation of pressure waves induced by the mutual interaction between the fluid and the compliant vessel walls. In particular, the interplay between anomalous pressure waves and pathologies like amputations, stenoses or devices like stents is of great interest from a medical viewpoint. However, the parameters that characterize reduced 1D models are often unknown, and feature variability not only from patient to patient, but also within the same individual, depending on physiological conditions (e.g., rest vs. stress, and young vs. old). This motivated the design of mathematical and numerical techniques to quantify the uncertainties in these models. Uncertainty Quantification (UQ) studies on the cardiovascular network entail two major challenges or limitations: (i) The employment of full 3D models for UQ analysis is extremely costly and requires computational resources that may not be easily accessible by users like hospitals, for financial, privacy or time constraints; (ii) Reduced 1D models may be inaccurate in capturing anomalies of the physiology in presence of cardiovascular pathologies like stenoses or aneurysms. Following the DDUQ approach (Carlberg et al, 2018), we enhance the efficiency and parallelism of the solvers by performing UQ at the subsystem level at each time step, and by propagating the information via Domain Decomposition techniques. We plan to enhance accuracy and reliability by replacing the 1D models with educated reduced models such as the Transversally Enriched Pipe Element Method (Mansilla et al, 2017), capable of retaining the local cross-sectional dynamics, approximately at the same cost of 1D reduced models.