- Record: found
- Abstract: found
- Article: found
CRISPR-Cas: Converting A Bacterial Defence Mechanism into A State-of-the-Art Genetic Manipulation Tool
Read this article at
Abstract
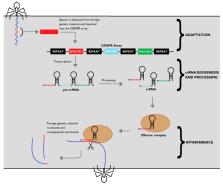
Bacteriophages are pervasive viruses that infect bacteria, relying on their genetic machinery to replicate. In order to protect themselves from this kind of invader, bacteria developed an ingenious adaptive defence system, clustered regularly interspaced short palindromic repeats (CRISPR). Researchers soon realised that a specific type of CRISPR system, CRISPR-Cas9, could be modified into a simple and efficient genetic engineering technology, with several improvements over currently used systems. This discovery set in motion a revolution in genetics, with new and improved CRISPR systems being used in plenty of in vitro and in vivo experiments in recent years. This review illustrates the mechanisms behind CRISPR-Cas systems as a means of bacterial immunity against phage invasion and how these systems were engineered to originate new genetic manipulation tools. Newfound CRISPR-Cas technologies and the up-and-coming applications of these systems on healthcare and other fields of science are also discussed.
Related collections
Most cited references111
- Record: found
- Abstract: found
- Article: not found
A TALE nuclease architecture for efficient genome editing.
- Record: found
- Abstract: found
- Article: not found
Targeting DNA double-strand breaks with TAL effector nucleases.
- Record: found
- Abstract: found
- Article: not found