- Record: found
- Abstract: found
- Article: not found
Myeloid-derived suppressor cells in COVID-19: A review
Read this article at
Abstract
Coronavirus disease 2019 (COVID-19) is a potentially life-threatening infection characterized by excessive inflammation, coagulation disorders and organ damage. A dysregulated myeloid cell compartment is one of the most striking immunopathologic signatures of this newly emerged infection. A growing number of studies are reporting on the expansion of myeloid cells with immunoregulatory activities in the periphery and airways of COVID-19 patients. These cells share phenotypic and functional similarities with myeloid-derived suppressor cells (MDSCs), which were first described in cancer patients. MDSCs are a heterogeneous population of pathologically activated myeloid cells that exert immunosuppressive activities against mainly effector T cells. The increased frequency of these cells in COVID-19 patients suggests that they are involved in immune regulation during this infection. In this article, we review the current findings on MDSCs in COVID-19 and discuss the complex role of these cells in the immunopathology of COVID-19.
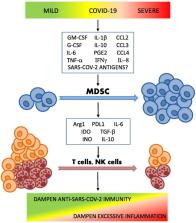
Graphical abstract
Related collections
Most cited references104
- Record: found
- Abstract: found
- Article: not found
SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor
- Record: found
- Abstract: found
- Article: found
COVID-19: consider cytokine storm syndromes and immunosuppression
- Record: found
- Abstract: found
- Article: not found

