- Record: found
- Abstract: found
- Article: found
Recent Advances in Detecting Mitochondrial DNA Heteroplasmic Variations
Read this article at
Abstract
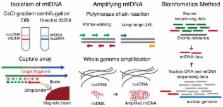
The co-existence of wild-type and mutated mitochondrial DNA (mtDNA) molecules termed heteroplasmy becomes a research hot point of mitochondria. In this review, we listed several methods of mtDNA heteroplasmy research, including the enrichment of mtDNA and the way of calling heteroplasmic variations. At the present, while calling the novel ultra-low level heteroplasmy, high-throughput sequencing method is dominant while the detection limit of recorded mutations is accurate to 0.01% using the other quantitative approaches. In the future, the studies of mtDNA heteroplasmy may pay more attention to the single-cell level and focus on the linkage of mutations.
Related collections
Most cited references74
- Record: found
- Abstract: found
- Article: not found
Detection of ultra-rare mutations by next-generation sequencing.
- Record: found
- Abstract: found
- Article: not found