- Record: found
- Abstract: found
- Article: found
Bacteria of the Genus Xenorhabdus, a Novel Source of Bioactive Compounds
Read this article at
Abstract
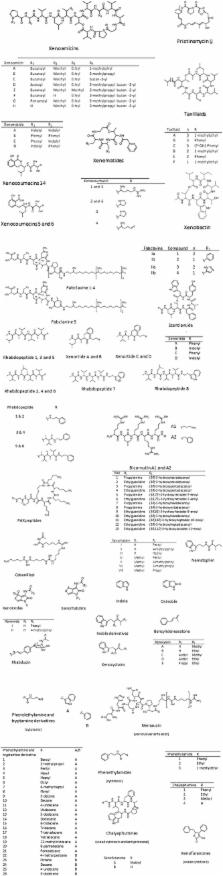
The genus Xenorhabdus of the family Enterobacteriaceae, are mutualistically associated with entomopathogenic nematodes of the genus Steinernema. Although most of the associations are species-specific, a specific Xenorhabdus sp. may infect more than one Steinernema sp. During the Xenorhabdus–Steinernema life cycle, insect larvae are infected and killed, while both mutualists produce bioactive compounds. These compounds act synergistically to ensure reproduction and proliferation of the nematodes and bacteria. A single strain of Xenorhabdus may produce a variety of antibacterial and antifungal compounds, some of which are also active against insects, nematodes, protozoa, and cancer cells. Antimicrobial compounds produced by Xenorhabdus spp. have not been researched to the same extent as other soil bacteria and they may hold the answer to novel antibacterial and antifungal compounds. This review summarizes the bioactive secondary metabolites produced by Xenorhabdus spp. and their application in disease control. Gene regulation and increasing the production of a few of these antimicrobial compounds are discussed. Aspects limiting future development of these novel bioactive compounds are also pointed out.
Related collections
Most cited references120
- Record: found
- Abstract: found
- Article: not found
Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study
- Record: found
- Abstract: found
- Article: not found
Waves of resistance: Staphylococcus aureus in the antibiotic era.
- Record: found
- Abstract: found
- Article: not found