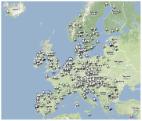
Introduction Staphylococcus aureus is the main cause of purulent infection in humans [1]. S. aureus has the potential for local as well as disseminated infection and can cause lesions in all tissues and anatomical sites. Infections can be either acquired in the community or in association with health care. The position of S. aureus as one of the most important human pathogens is largely due to its virulence potential and ubiquitous occurrence as a coloniser in humans, domestic animals, and livestock [2]. Between 25% and 35% of healthy human individuals carry S. aureus on the skin or mucous membranes [3]. Any injury that compromises epithelial integrity, trauma, medical or surgical interventions, as well as viral infections, can lead to tissue invasion [4]–[6]. It is assumed that severity and outcome depend largely on the virulence of the introduced strain and the immune repertoire of the host [7],[8]. Occasionally, S. aureus acquires enhanced virulence and antimicrobial resistance through horizontal DNA transfer and maintains these mobile genetic elements in a predominantly clonal genomic background. Thus, clones of S. aureus are relatively stable and mainly diversify by the accumulation of single nucleotide substitutions in the absence of frequent interstrain recombination. It is therefore possible to discern different clones and clonal lineages by molecular typing [9]. This method allows several important observations to be made regarding the evolution, epidemiology, and spread of clones with particular public health importance, such as hospital-, community-, and livestock-associated methicillin-resistant S. aureus (MRSA). For MRSA, this surveillance is particularly important because it appears that certain clones have disseminated over wide geographical regions and are threatening major improvements in curative and public health medicine [10]. A geographically detailed description of this expansion on a continent-wide scale has been inadequate, however, due to the lack of appropriate surveys and agreement on a consistent application of standardized molecular typing approaches. At the same time, little is known about the population structure and geographical abundance of methicillin-susceptible S. aureus (MSSA), which provides the genetic reservoir from which MRSA emerge. The present study was designed to fill these knowledge gaps and to provide (i) the first representative and contemporaneous snapshot of the genetic population structure of S. aureus (based on spa typing) that cause invasive infection in the European region; (ii) insights into the geographic distribution of different clones among MSSA and MRSA on a continent-wide scale; and (iii) a public Web-based mapping tool supplying interactive access and an intuitive illustration of the results generated by this large-scale typing initiative. The current study was also set up to establish the logistics for future collaborative studies that will continue to improve crucial knowledge for clinicians and diagnostic laboratories about the geographic and temporal dynamics of the MSSA/MRSA clones and their epidemic patterns in neighbouring geographical areas. Lastly, the study is intended to additionally strengthen the role of the S. aureus Reference Laboratories (SRLs) by exposing and communicating potentially important public health threats to health professionals and the general public. Methods spa Typing Epidemiological typing uses highly discriminatory genetic markers that characterize human pathogens allowing the identification of isolates that are closely related due to recent common ancestry. The spa locus of S. aureus codes for protein A, a species-specific gene product known for its IgG binding capacity. This locus is highly polymorphic due to an internal variable region of short tandem repeats, which vary not only in numbers but also because of nucleotide substitutions within individual repeat units [11]. DNA sequences of the spa gene therefore provide portable, unambiguous, and biologically meaningful molecular typing data, which have demonstrated their utility for epidemiological purposes such as transmission and outbreak investigations at various geographical levels [12],[13]. Capacity Building During three workshops organised for technical personnel from 28 European SRLs, participants received hands-on training in spa sequence typing and spa sequence analysis according to a standard protocol using a purpose-designed software tool, StaphType, developed by Ridom GmbH [13]. Proficiency testing was carried out by mailing each SRL five well-characterized S. aureus isolates and five sequence chromatograms (trace files) of known spa types as described previously [14],[15]. All laboratories participating in the structured survey described here fulfilled quantifiable quality criteria, which consisted of an unambiguous base-calling of all sequenced nucleotides for both forward and reverse sequencing runs of the test panel. Structured Survey A protocol was drawn up and circulated for comment to all SRLs and agreed upon in April 2006. Following this protocol, European SRLs were asked to identify approximately 20 laboratories that serve hospitals and that are geographically and demographically representative of their country, and secure their participation. These laboratories were chosen from those that regularly participate in the European Antimicrobial Resistance Surveillance System (EARSS). For 6 mo, from September 2006 until February 2007, these participants were asked to submit the first five successive MSSA isolates and the first five successive MRSA isolates from individual patients with invasive infection. An invasive infection was defined as a localised or systemic inflammatory response to the presence of S. aureus at otherwise sterile anatomical sites. Isolates were dispatched by the participating laboratories to the SRLs accompanied by additional information, including the EARSS laboratory identifier, the sample number, the date of isolation, origin of clinical specimen, demographic detail (such as age and gender), epidemiological context (hospital-acquired when symptoms developed more than 48 h after admission or as community-onset otherwise), isoxazolylpenicillin- (i.e., oxacillin) or cefoxitin-resistance, and all-cause mortality 14 d after isolation of the first invasive isolate. SRLs confirmed MRSA by mecA PCR or determination of minimum inhibitory concentration for oxacillin together with PBP2a agglutination. Additional data were uploaded to the database and Web application if available. These consisted of staphylococcal cassette chromosome mec (SCCmec) typing, and identification of luk-PV genes, indicative of Panton-Valentine Leukocidin (PVL). All SRLs preserved the isolates in strain collections and performed spa sequence typing according to the standard protocol, uploaded the sequence information, and made this available by synchronisation with the central Ridom SpaServer (www.spaserver.ridom.de) curated by SeqNet.org at the Institute of Hygiene, University Hospital Münster, Germany [15],[16]. Submitted sequences were quality controlled by comparison with accompanying chromatograms (trace files) and excluded if stringent quality criteria for excellent sequencing data were not fulfilled. spa types were grouped into spa complexes if a single genetic event such as single insertions, single deletions, or single nucleotide polymorphism could account for the observed sequence divergence. In the following the designation of spa types indicated by lower case “t” follow the nomenclature used by the spa server and multilocus sequence types are reported as sequence type (ST) according to convention [17]. Finally, the SCCmec type is also added to a string consisting of spa type/ST/SCCmec all in parenthesis, e.g., (t032/ST22/SCCmecIV). Epidemiological and typing data were communicated in parallel to a central SQL database at the National Institute for Public Health and the Environment (RIVM) of the Netherlands. For each participating laboratory, SRLs also provided the postal address and indicated their administrative region (such as region, province, state, department, or NUTS-2 level) if the location of laboratories were to be aggregated on a higher administrative geographical level for display on the interactive mapping tool (which was done only for Austria, Belgium, Czech Republic, and Poland). All data were anonymous and collected in accordance with the European Parliament and Council decision for the epidemiological surveillance and control of communicable disease in the European community [18],[19]. Ethical approval and informed consent were thus not required. Data Analysis, Geographical Illustration, and Cluster Identification All data were inspected for inconsistencies and analysed on a country-by-country basis and returned to SRLs for feedback, clarification of inconsistencies and final approval in February 2008. After final approval, data were analysed using Stata version 10 and SAS version 9.1 (SAS Institute Inc.) using chi-square test for proportions and Student's t-test for continuous variables. The index of diversity is an unbiased measure of the probability of drawing two different spa types given the distribution of spa types in the sample. The 95% confidence intervals (CIs) were calculated as described previously [20]. Postal address and location of all sampling laboratories were converted into decimal cartesian coordinates using the geocoding facility at www.spatialepidemiology.net [21]. Pairwise distances of laboratories that isolated MSSA and MRSA with identical spa types were calculated and distance matrices were summarised and compared by nonparametric tests. The Web application SRL-Maps (http://www.spatialepidemiology.net/srl-maps) was developed to interrogate the data on the basis of mapping of laboratory locations and on centroids of administrative regions (when laboratory results were aggregated at the level of administrative region). A spatial scan statistic was used to identify the geographic distribution of specific spa types at two levels: (i) country-specific frequencies that take into account all spa types within national boundaries and (ii) regional clusters of varying size independent of national boundaries. To identify spa types with higher than expected occurrence in any of the participating countries, the observed number of all spa types isolated within each country was compared with the number expected under the assumption of a European-wide random distribution. For the identification of regional clusters, circular windows around all laboratory locations were projected. For each location, the radius of the window was varied from 0 to 1,000 km. In this way, a finite number of distinct windows was created. For each window, the observed number of isolates with a specific spa type was compared with the expected number under the assumption of a random distribution. 10,000 random distributions were obtained by varying the composition of local spa types at all laboratory locations consistent with cumulative spa-specific frequencies using Monte Carlo Simulations. The test statistic was calculated by log-likelihood ratio test, whereby countries or windows where the observed spa type frequencies differed from those obtained by simulation were considered to contain significant clusters with an alpha error of p<0.0001. The scan-statistic was executed with SaTScan software using SAS macros [22],[23]. Results Summary Statistics 26 SRLs from 26 countries satisfactorily fulfilled the proficiency criteria for spa sequence typing and contributed data for final analysis. Between September 2006 and February 2007, 357 laboratories serving 450 hospitals (Figure 1) collected 2,890 successive MSSA and MRSA isolates from patients with invasive S. aureus infection (2,603 from blood cultures, 90.1%; 17 from cerebrospinal fluid, 0.6%; and 270 from puncture fluids of other normally sterile anatomical sites, 9.3%). Final inspection of data revealed missing information for gender (28 isolates, 1%), age (54 isolates, 1.9%), sampling dates (74 isolates, 2.6%), epidemiological context (community onset or hospital acquisition, 568 isolates 19.7%), all-cause mortality 14 d after S. aureus isolation (1052 isolates, 36.4%), and spa type (40 isolates, 1.4%). Table 1 gives a summary overview of the number of participating laboratories, isolates, and spa types submitted by country. Many laboratories were unable to collect five invasive MRSA isolates within the sampling period because of a low MRSA incidence in the hospitals they serve. Therefore, the combined collection consisted of two-thirds MSSA (1,923; 66.5%) and one-third MRSA (967; 33.5%, Table 2). Patients with invasive MRSA infections were older (Figure 2) with a median age of 69 y compared to a median age of 63 y in MSSA patients (p<0.001). Moreover, MRSA patients had higher all-cause mortality (20.8%) 14 d after the first isolation of S. aureus than MSSA patients (13.2%, p<0.0001). More males (1,765/2,862; 61.7%) than females had invasive S. aureus infections. The proportion of MRSA compared to MSSA did not differ between the sexes (p = 0.2). Of the 231 MRSA that were reported as community-onset (CO-MRSA), 226 (97.8%) were tested for the presence of PVL-associated genes luk-PV and ten (4.4%) were positive. Of the 585 hospital-acquired MRSA (HA-MRSA), 551 (94.2%) were tested for PVL and six (1.1%) were positive. The difference was significant (p = 0.009). 10.1371/journal.pmed.1000215.g001 Figure 1 Locations of participating laboratories. 10.1371/journal.pmed.1000215.g002 Figure 2 Age distribution and all-cause mortality of patients 14 d after diagnosis of invasive S. aureus infections in Europe. Age divided in bands of 5 y, except for infants under 1 y (−1). 10.1371/journal.pmed.1000215.t001 Table 1 Summary overview of participating laboratories, hospitals, number of invasive isolates of MSSA and MRSA, and spa types by country. Country n Labs n Hospitals n Isolates MSSA MRSAa n spa Types MSSA n spa Types MRSA n Not Typeable Percent Not Typed Austria 18 48 174 120 54 70 19 1 0.6 Belgium 22 22 195 107 88 65 25 1 0.5 Bulgaria 8 8 54 29 25 23 11 0 0.0 Croatia 11 11 88 50 38 27 13 6 6.8 Cyprus 1 1 16 9 7 8 5 0 0.0 Czech Republic 20 20 145 94 51 64 9 0 0.0 Denmark 14 30 112 108 4 72 2 0 0.0 Finland 5 5 22 15 7 14 7 0 0.0 France 23 23 225 114 111 75 27 0 0.0 Germany 27 27 180 98 82 56 20 1 0.6 Greece 3 3 35 20 15 12 6 6 17.1 Hungary 10 13 110 66 44 35 9 2 1.8 Iceland 1 1 5 5 0 5 0 0 0.0 Ireland 22 22 169 85 84 55 26 0 0.0 Italy 19 19 147 80 67 53 15 0 0.0 Latvia 11 12 43 38 5 20 1 0 0.0 Malta 1 1 15 3 12 2 5 3 20.0 Netherlands 18 21 204 195 9 98 9 6 2.9 Norway 11 20 55 55 0 38 0 1 1.8 Poland 23 23 179 132 47 42 14 0 0.0 Portugal 12 12 88 48 40 36 13 0 0.0 Romania 10 10 36 25 11 18 3 0 0.0 Slovenia 11 12 58 48 10 29 3 2 3.4 Spain 21 21 204 113 91 58 19 1 0.9 Sweden 20 47 200 195 5 90 5 3 1.5 UK 15 18 131 71 60 51 18 7 5.3 Total 357 450 2,890 1,923 967 565 155 40 1.4 a The number of MRSA isolates does not reflect a prevalence or occurrence in particular countries as the protocol requested submission of the first five MSSA and MRSA isolates. 10.1371/journal.pmed.1000215.t002 Table 2 Summary statistics of S. aureus isolated in 26 European countries. Statistics n a MSSA MRSA Total/Overall p-Value* Frequency (%) 2,890 1,923 (66.5) 967 (33.5) 2,890 (100%) — Median age (IQR) 2,836 63 (46–75) 69 (55–78) 66 (49–76) <0.0001 Male gender (%) 2,862 1,159 (60.8) 606 (63.3) 1,765 (61.7) 0.2 All-cause mortality after 14 d (%) 1,838 153 (13.2) 141(20.8) 294 (16.0) <0.0001 Hospital acquisition (%) 2,322 777 (51.6) 585 (71.7) 1,362 (58.7) <0.0001 N spa types 2,850 565 155 660b — N not typeable 2,850 27 (1.4) 13 (1.3) 40 (1.4) 0.9 Index of diversity (95% CI) 2,850 0.985 (0.983–0.987) 0.940 (0.933–0.947) 0.983 (0.982–0.984) <0.05c Mean distance in kilometres between laboratories that isolated identical spa types (95% CI) 1,614d 1,046.2 (1109.5–983.0) 786.8 (975.7–597.9) — 0.03d * p-Value for the comparison of MSSA versus MRSA. a Number of isolates for which data were available. b Total number of spa types includes 60 spa types that contain both MSSA and MRSA. c Deduced from non-overlapping 95% confidence intervals. d Includes only MRSA and MSSA with more than ten isolates per spa type. IQR, interquartile range. spa Typing A total of 660 spa types were reported (Table 2). Among all spa types, 565 and 155 were assigned to MSSA and MRSA, respectively, of which, 505 were exclusively identified as MSSA and 95 for MRSA alone. 60 spa types contained both MSSA and MRSA. 27 of the MSSA (1.4%) and 13 of the MRSA (1.3%) isolates were nontypeable. Table 3 shows the rank order of the most frequent spa types isolated during the survey and Table 4 the three most frequent spa types by country. MRSA was less diverse than MSSA. Only five spa types accounted for almost half (48.1%) of all MRSA isolates, whereas the same proportion of MSSA isolates comprised the 26 most frequent MSSA spa types. Estimates of the genetic diversity differed significantly with an index of diversity for MSSA of 0.985 (95% CI 0.983–0.987) and 0.940 (95% CI 0.933–0.947) for MRSA. While MSSA diversity ranged between 0.934 in Latvia and 1.0 in Iceland (unpublished data), MRSA diversity between countries was more heterogeneous ranging from 0.62 in Romania to 0.91 in Austria (Figure 3), indicating the presence of few dominant MRSA spa types in several countries. Accordingly, MRSA showed a higher degree of geographic clustering as the average distance between laboratories that isolated the same spa type was significantly smaller than for MSSA (Table 2). No correlation between genetic diversity of MRSA and overall proportion of MRSA among S. aureus blood stream infections at country level as reported to the EARSS database for 2007 was found (r = −0.09, p = 0.75) [24], indicating that single successful spa types cannot explain the variance in the proportion of MRSA causing S. aureus blood stream infections observed in Europe. 10.1371/journal.pmed.1000215.g003 Figure 3 Estimates of country-specific genetic diversity expressed as Simpson's index of diversity of spa types (as a percentage) for MSSA (light blue diamonds) and MRSA (dark blue diamonds) and 95% CIs (bars). Only countries for which spa type information for more then ten MRSA isolates were available were included in this figure. 10.1371/journal.pmed.1000215.t003 Table 3 20 most frequent spa types and their STs among MSSA and MRSA isolated in 26 European countries. Rank MSSA Frequency Percent Cumulative Percent Rank MRSA Frequency Percent Cumulative Percent spa Type MLST spa Type MLST 1 t002 ST-5a, S-231a 93 4.8 4.8 1 t032 ST-22a 140 14.5 14.4 2 t084 ST-15a (ST-18) 89 4.6 9.5 2 t008 ST-8a (ST-247, ST-250, ST-254) 120 12.4 26.8 3 t015 ST-45a 84 4.4 13.8 3 t041 ST-111a, ST-228a 72 7.4 34.2 4 t091 ST-7a 82 4.3 18.1 4 t003 (ST-5) ST-225a 71 7.3 41.6 5 t012 ST-30a 77 4.0 22.1 5 t002 ST-5a, ST-231a 62 6.4 48.1 6 t127 ST-1a 57 3.0 25.1 6 t067 ST-5a, ST-125a 50 5.2 53.3 7 t008 ST-8a (ST-247, ST-250, ST-254) 55 2.9 27.9 7 t001 (ST-5, ST-222) ST-228a 30 3.1 56.4 8 t021 ST-30a (ST-33, ST-55) 49 2.5 30.5 8 t037 ST-239a (ST-240), ST-241a 27 2.8 59.2 9 t005 ST-22a (ST-23, ST-60) 42 2.2 32.7 9 t030 ST-239a (ST-246) 20 2.1 61.2 10 t026 (ST-45, ST-47) 27 1.4 34.1 10 t024 ST-8a 14 1.4 62.7 11 t065 (ST-45, ST-46) 26 1.4 35.4 11 t190 ST-8a 14 1.4 64.1 12 t160 (ST-12, ST-13) 26 1.4 36.8 12 t515 ST-22a 12 1.3 65.5 13 t056 (ST-101) 25 1.3 38.1 13 t038 ST-45a 12 1.2 66.7 14 t050 ST-45a 21 1.1 39.2 14 t022 ST-22a 11 1.1 67.8 15 t078 (ST-26) 21 1.1 40.2 15 t740 ST-45a 11 1.1 69.0 16 t164 (ST-20) 19 1.0 41.2 16 t012 ST-30a 9 0.9 69.9 17 t346 (ST-15, ST-620) 18 0.9 42.2 17 t015 ST-45a 9 0.9 70.8 18 t024 ST-8a 17 0.9 43.1 18 t044 ST-80a 9 0.9 71.8 19 t230 ST-45a 17 0.9 43.9 19 t045 ST-5a (ST-225) 8 0.8 72.6 20 t166 (ST-34) 16 0.8 44.8 20 t127 ST-1a 8 0.8 73.4 — Other — 1,062 55.2 100.0 — Other — 258 26.6 100.0 Total — — 1,923 100 — — — — 967 100 — STs in parentheses are those associated with the spa type in the SeqNet.org Spa typing data base. a MLST as determined by SRLs. 10.1371/journal.pmed.1000215.t004 Table 4 First, second, and third most frequent MSSA and MRSA spa types per country and their relative proportions. Country MSSA spa Type MRSA spa Type n 1st (%) 2nd (%) 3rd (%) n 1st (%) 2nd (%) 3rd (%) Austria 120 t091 (8.3) t002 (6.7) t012 (5.0) 54 t190 (18.5) t041 (16.7) t001 (14.8) Belgium 107 t002 (9.4) t209 (4.7) t012, t091, t740 (3.7) 88 t008 (29.6) t002, t038 (13.7) t740 (12.5) Bulgaria 29 t056 (10.3) t078, t148, t156, t1346 (6.9) a 25 t030 (36.0) t037 (16.0) t010 (12.0) Croatia 50 t050 (10.0) t005, t015, t1361 (8.0) t164 (6.4) 38 t041 (36.8) t091 (10.5) t026, t1003 (8.3) Cyprus 9 t002 (22.2) a — 7 t012, t30 (28.6) a — Czech Republic 94 t015, t130 (5.3) t024, t122, t1081 (4.3) t056, t156, t491, t1231 (3.2) 51 t003 (66.7) t032 (13.7) t002 (7.8) Denmark 108 t230 (7.4) t002, t127 (4.6) t065, t084 (3.7) 4 t024, t037 (50.0) — — Finland 15 t026 (13.3) a — 7 b — — France 114 t002 (11.4) t008 (6.1) t012 (4.4) 111 t008 (48.6) t777 (7.2) t024 (5.4) Germany 98 t008 (11.2) t084 (7.1) t015, t091 (6.1) 82 t032 (35.4) t003 (28.1) t001 (8.6) Greece 20 t267 (15.0) t012 (10.0) a 15 t002, t044 (26.7) t037 (20.0) a Hungary 66 t091, t216 (10.6) t012, t084 (7.60) t002, t015, t2115 (4.7) 44 t032 (38.6) t041 (25.0) t062 (13.6) Iceland 5 b — — 0 — — — Ireland 85 t021 (7.1) t012 (4.7) t078, t127, t166, t382, t548 (3.6) 84 t032 (45.2) t515 (9.5) t022 (4.8) Italy 80 t091 (10.04) t084 (8.8) t012 (7.5) 67 t041 (34.3) t008 (28.4) t001 (13.4) Latvia 38 t435 (21.1) t015 (13.2) t313, t698 (7.9) 5 b — — Malta 3 b — — 12 t001, t032 (30.0) t012 (20.0) t002, t022 (10.0) Netherlands 195 t091 (7.7) t127 (6.2) t002, t012, t084 (4.2) 9 b — — Norway 55 t065 (9.1) t084 (9.1) t002, t015, t095 (3.7) 0 — — — Poland 132 t127 (12.9) t084 (9.9) t015 (7.6) 47 t037 (29.81) t003, t015 (14.9) t002, t041, t1574 (6.4) Portugal 48 t008 (8.3) t002, t645 (6.3) t021, t127, t148, t148, t179, t189 (4.2) 40 t032 (32.5) t002 (20.0) t535, t747, t2357 (7.5) Romania 25 t021, t284 (12.0) t005, t008, t450 (8.0) a 11 t030 (54.6) t127 (36.4) t015 (9.1) Slovenia 48 t091 (20.8) t015 (10.4) t005 (8.5) 10 t041 (70.0) a — Spain 113 t002 (12.4) t012, t067 (8.0) t015 (4.5) 91 t067 (47.3) t002 (15.4) t008 (7.7) Sweden 195 t015 (9.2) t084 (8.2) t012 (5.7) 5 b — — UK 71 t012, t127 (5.6) t021 (4.2) nine different types 60 t032 (61.7) t788, t1516 (3.3) a a All remaining spa types equally distributed. b No ranking, all spa types equally distributed. Clustering of spa Types at Country and Regional Level In 17 countries, 22 spa types were found with frequencies that were unexpected when applying the hypothesis of a random distribution, indicative of local epidemics (Table 5). Most (86.9%) of these were MRSA. In ten countries two spa types coexisted with unexpected frequencies. In four of them, these two spa types showed a close genetic relationship and belonged to the same spa complex whereby a single genetic event could account for the sequence divergence between the types. There was also a frequent regional coincidence with neighbouring countries sharing identical epidemic spa types. The Czech Republic and Germany shared spa type t003 (t003/ST225/SCCmecII), Bulgaria and Romania shared t030 (t030/ST239/SCCmecIII), the UK and Ireland t032 (t032/ST22/SCCmecIV), Italy and Croatia shared t041 (t041/ST228/SCCmecI), and strain t067 (t067/ST5125/SCCmecIV) whose dominance in Spain was first identified through this initiative [25], was also found in southern France. The notion of regional spread was supported by the cluster statistic that projected windows beyond national boundaries for this dataset (Table 6). The degree of unexpectedness, which is an indication of the significance of each cluster, is expressed by the log likelihood ratio (LLR). The majority of regional clusters extended beyond national boundaries and 74% of all isolates that occurred in these clusters were MRSA. The most significant cluster was identified in Spain and consisted of spa type t067 (t067/ST5&125/SCCmecIV). A northern Balkan/Adriatic cluster consisting of spa type t041 (t041/ST228/SCCmecI) was found in Austria, Hungary, Slovenia, Croatia, and northern and central Italy. In Britain and Ireland, t032 (t032/ST22/SCCmecIV), known as epidemic MRSA 15 (EMRSA-15), was the dominant type and represented the third most significant cluster. An additional cluster of spa type t032, albeit less significant and much smaller, was located in the Brandenburg area of Germany. Central Germany, the Czech Republic, and western Poland were included in a large regional cluster of spa type t003 (t003/ST225/SCCmecII), which was geographically centred in Saxony and had a radius of approximately 400 km corresponding to the German hospitals participating in the study. Figure 4 provides a geographical illustration of these clusters. The largest cluster in size as well as in numbers (radius 930 km, 119 isolates) consisted of spa type t008 and was centred in southern France. This cluster consisted of ST8 and contained different subclones as it included both MSSA and MRSA, and MRSA isolates exhibited two different SCCmec elements (SCCmecI and IV). A smaller cluster, ranking in sixth position in terms of significance, was located in Flanders on the Belgian-Dutch border and consisted of spa type t740 (t740/ST45/SCCmecIV). Interestingly, regional spa clusters with overlapping geographical range were frequently made up of spa types that belonged to the same spa complex, a clear indication that local spread is accompanied by local evolution of the rapidly evolving spa locus. Clusters with the smallest size (0 km) included those submitted by single laboratories most likely reflecting single hospital outbreaks. Three regional clusters consisted of MSSA alone. They were located in Latvia (t435/ST425), Poland (t127/ST1), and Denmark (t230/ST45), indicating that regional spread of S. aureus is not limited to MRSA alone. 10.1371/journal.pmed.1000215.g004 Figure 4 Location of laboratories isolating S. aureus of spa types t067, t041, t032, and t003, which are the four most significant regional clusters on SRL-Maps. The numbers within each placemark represent the number of isolates and the colours represent resistance phenotypes: red, MRSA; green, MSSA; yellow, a mixture of MRSA and MSSA. 10.1371/journal.pmed.1000215.t005 Table 5 Unexpectedly frequent spa types at country level assuming a European-wide random distribution. Country Cluster Number Country spa Type spa Complexa STb n Labs Reporting Clustered spa Type n Labs Participating in Survey Percent Labs Reporting Clustered spa Type n Isolates Observed n Isolates Expectedc n MRSA among Observed Isolates Percent MRSA among Observed Isolates 1 Austria t190 190 8 7 18 38.9 11 0.9 10 91 2 Belgium t740 740 45 6 22 27.3 15 1.0 11 73 3 Belgium t038 740 45 8 22 36.4 12 0.8 12 100 4 Bulgaria t030 12 239 4 8 50.0 10 0.4 9 90 5 Croatia t041 1 228 7 11 63.6 14 2.2 14 100 6 Czech Republic t003 45 225 16 20 80.0 34 3.8 34 100 7 Czech Republic t130 130 — 3 20 15.0 5 0.4 5 100 8 Denmark t230 728 45 7 14 50.0 8 0.7 0 0 9 France t008 8 8 22 23 95.7 61 13.6 54 89 10 France t777 777 5 6 23 26.1 9 0.7 8 89 11 Germany t003 45 225 10 27 37.0 24 4.7 23 96 12 Germany t032 32 22 9 27 33.3 29 9.5 29 100 13 Greece t044 44 80 3 3 100.0 5 0.1 4 80 14 Hungary t062 Singleton 5 2 10 20.0 6 0.3 6 100 15 Hungary t216 Singleton 59 7 10 70.0 7 0.5 7 100 16 Ireland t032 32 22 18 22 81.8 38 8.9 38 100 17 Ireland t515 32 22 7 22 31.8 8 0.7 8 100 18 Italy t041 1 228 13 19 68.4 23 3.7 23 100 19 Italy t001 1 228 8 19 42.1 9 1.5 9 100 20 Latvia t435 435 427 4 11 36.4 8 0.2 8 100 21 Latvia t425 425 368 3 11 27.3 5 0.1 5 100 22 Poland t037 12 239 11 23 47.8 21 2.2 14 67 23 Poland t127 127 1 6 23 26.1 17 4.0 17 100 24 Romania t030 12 239 3 10 30.0 6 0.3 6 100 25 Spain t067 2 5 & 125 18 21 85.7 52 4.4 43 81 26 Spain t002 2 5 15 21 71.4 28 10.9 14 50 27 United Kingdom t032 32 22 12 15 80.0 39 6.9 27 96 Total — — — — 235 — — 504 83.4 438 87 a spa complexes group spa types that differ by only a single evolutionary event (single indel or nucleotide polymorphism) into the same complex. b As determined by SRLs. c Average number of isolates with this spa type expected in country after 10,000 simulations on the basis of European-wide cumulative frequency. 10.1371/journal.pmed.1000215.t006 Table 6 Regional clusters of spa types. Regional Cluster Number spa Type spa Complexa STb Window Centre Window Radius (km) Countries reporting Clustered spa Type within Window n Isolates Observed n Isolates Expectedc Log Likelihood Ratio n MRSA among Observed Isolates Percent MRSA among Observed Isolates 1 t067 2 5 & 125 Alicante, Spain 716 ES, FR 55 4.7 126.9 46 84 2 t041 1 228 Split, Croatia 522 AT, HR, HU, SI, IT 59 10.6 84.76 59 100 3 t032 32 22 Belfast, Northern Ireland (UK) 596 IE, UK 77 15.9 84.74 75 97 4 t003 45 225 Leipzig, Germany 386 CZ, DE, PL 58 10.0 82.26 54 93 5 t008 8 8 Perpignan, France 931 AT, BE, DE, ES, FR, HR, PT, SI 119 45.5 72.96 105 88 6 t740 740 45 Goes, Netherlands 81 BE, NL 15 0.6 50.36 11 73 7 t030 12 239 Pleven, Bulgaria 331 BG, RO 16 0.7 45.24 15 94 8 t037 12 239 Plock, Poland 330 PL 21 2.2 36.02 18 86 9 t038 740 45 Wilrijk, Belgium 92 BE 12 0.8 32.71 12 100 10 t190 190 8 St Pölten, Austria 56 AT 10 0.3 29.63 9 90 11 t001 1 228 Sibenik, Croatia 885 AT, DE, HU, IT, MT 29 10.2 27.55 29 100 12 t435 435 427 Daugavpils, Latvia 189 LV 8 0.2 25.82 0 0 13 t425 239 368 Riga, Latvia 0 LV 5 0.1 22.76 5 100 14 t127 127 1 Inowroclaw, Poland 131 PL 12 0.9 22.65 0 0 15 t032 32 22 Berlin, Germany 128 DE 19 3.2 22.2 19 100 16 t015 15 45 Lomza, Poland 581 CZ, LV, PL 35 11 20.72 7 20 17 t091 91 7 Ried, Austria 771 AT, BE, CZ, DE, FR, HR, HU, IT, NL, PL, SI 71 42.8 20.68 4 6 18 t777 777 5 Laval, France 277 FR 7 0.3 20.61 7 100 19 t081 78 25 Maków Mazowiecki, Poland 0 PL 4 0.0 19.37 3 75 20 t515 32 22 Mullingar, Ireland 196 IE, Northern Ireland (UK) 9 0.7 18.94 9 100 21 t002 2 5, 231 Coimbra, Portugal 484 ES, PT 33 10.6 18.37 20 61 22 t230 728 45 Hilleroed, Denmark 235 DE, DK, SE 11 1.4 17.19 0 0 23 t044 44 80 Nicosia, Cyprus 916 GR, CY 6 0.2 17.09 6 100 24 t2054 8 8 St Mande, France 226 BE, FR4 5 0.2 16.64 3 60 25 t062 singleton 5 Szolnok, Hungary 145 HU 6 0.3 15.24 6 100 Total — — — — 368 (mean) 339 702 173.1 — 522 74 Analyses were performed for spa types with five or more isolates during the study period. a spa complexes group spa types that differ by only a single evolutionary event (single indel or nucleotide polymorphism) into the same complex. b As determined by SRLs. c Average number of isolates with this spa type expected in window after 10,000 simulations on the basis of European-wide cumulative frequency. AT, Austria; BE, Belgium; BG, Bulgaria; CY, Cyprus; CZ, Czech Republic; DE, Germany; DK, Denmark; ES, Spain; FI, Finland; FR, France; GR, Greece; HR, Croatia; HU, Hungary; IE, Ireland; IT, Italy; LV, Latvia; MT, Malta; NL, Netherlands; PL, Poland; PT, Portugal; RO, Romania; SE, Sweden, SI, Slovenia; UK, United Kingdom. SRL-Maps The Web application SRL-Maps (http://www.spatialepidemiology.net/srl-maps) provides a community tool for the interrogation of the geographic distribution of different spa types. All laboratory and regional locations across Europe are represented as placemarks on a Google map. Clicking on a placemark displays, below the map, all spa types identified at that location (and their frequency) along with the number of isolates (and number of geographic locations) found elsewhere (if any) for each of these spa types (Figure 5). The European distribution of any spa type can then be displayed and placemarks on the map are colour-coded on the basis of whether the isolates at each location are all MSSA (green), all MRSA (red), or are a mix of MSSA and MRSA (yellow), with the number of isolates inside the placemark. Graphical charts are displayed showing spa type-specific proportion of MRSA/MSSA, all-cause mortality at 14 d, and the age distribution among cases (Figure 5). This method allows the identification and mapping of strains with particular public health importance and further exploration by the scientific community is encouraged. 10.1371/journal.pmed.1000215.g005 Figure 5 Location of laboratories isolating S. aureus spa type t067 (as shown in Figure 4) viewed using SRL-Maps. Isolates from LAB ES056 in Madrid, Spain, and the distribution of all other t067 isolates are shown (n = 62). Each placemark indicates whether isolates are MSSA (green), MRSA (red), or a mixture (yellow) and the selected laboratory is blue. The numbers of isolates are indicated inside the placemark. The pie charts on the right show the proportion of MRSA/MSSA and all-cause mortality after 14 d, and the bar chart displays patient age distribution. Observations on Specific spa Types For both MSSA and MRSA isolates, there was no association between specific spa types and all-cause mortality after 14 d, indicating that no spa type was associated with hyper-virulence. Of the ten CO-MRSA isolates that were found to be PVL-positive, three were assigned to spa type t044 (t044/ST80/SCCmecIV), the so-called European community-acquired (CA)–MRSA, and another three had spa type t008 (t008/ST8/SCCmecIV) and are indistinguishable from USA300 CA-MRSA. Of the four other PVL-positive CO-MRSA, two belonged to t622 (spa complex 8/ST8/SCCmecIV), one to t529 (ST1043/SCCmecV), and one to t437 (not further characterised). MRSA belonging to ST398 have recently emerged in several European countries and are regarded as being livestock-associated (LA-MRSA). Of all spa types typically associated with this clone, spa types t011, t034, t571, t1255, and t2383 were identified on 12 occasions (1.3%) in eight countries during this survey. None of these isolates, however, displayed an MRSA phenotype or harboured the mecA gene. Discussion Predominant spa types showed a wide geographic distribution. The average distance between the locations from which the same spa types were sampled was smaller for MRSA isolates than MSSA isolates, suggesting a higher degree of geographical clustering of MRSA. Moreover, genetic diversity was much lower for invasive MRSA than MSSA and differed considerably between countries. Spatial-scan statistics corroborated a fundamental difference between MRSA and MSSA with respect to regional dissemination. The majority of isolates that formed regional clusters were MRSA, and 13 of the 15 major MRSA spa types (defined as more than ten isolates in the database) occurred in geographical clusters. They were typically hospital acquired and no more than three clusters overlapped in the same region. Conversely, of the 27 major MSSA spa types, only five showed significant geographical clustering and only three consisted of MSSA alone. Thus, invasive MRSA clones in Europe display a typical epidemic behaviour and have a predominantly regional distribution. To unravel the dynamics of spread of these epidemic MRSA requires the present type of survey to be repeated every few years. The emergence of MRSA occurs by the acquisition of the methicillin resistance determinant (SSCmec) by MSSA strains. MRSA strains typically emerge from the most prevalent MSSA strains and it is a rare event, although new findings suggest that it is more frequent than originally suggested [26],[27]. There are thus fewer MRSA clones compared to MSSA clones and they are very young on evolutionary time scales (less than 50 y old) and have had little time to diversify since they arose, whereas MSSA are much older and thus more diverse. MRSA clones also expand because of the selective forces exerted by heavy antibiotic use in hospitals and conditions that favour transmission within and between hospitals, which constrains their diversity. In contrast, MSSA will be subject to much weaker selection leading to neutral genetic drift that maintains their diversity. Geographic spread of MRSA clones will be facilitated by repeated hospital admissions and referrals of MRSA carriers [28] who typically belong to an older and thus less mobile segment of the population. The broader distribution of MSSA clones may reflect dissemination by hosts with different travel patterns than MRSA carriers as well as the longer time that MSSA clones have had to spread. The present survey set several precedents in the field of molecular epidemiology of S. aureus. First, it brought together reference laboratories from 26 European countries adopting a standardised quality-controlled DNA sequence-based typing method; second, it provided a contemporaneous and comprehensive population snapshot of S. aureus isolates from invasive disease using an agreed sampling frame; third, it utilised modern spatial scan statistics to identify geographical clustering; fourth, it provided the first public domain Web-based interactive mapping tool for future public health research; and finally, it consolidated a collaborative framework for the continuation of this important European surveillance initiative. All Member States of the European Union except Estonia, Lithuania, Luxemburg, and Slovakia participated and variously achieved a country-wide enrolment of diagnostic laboratories. In the run-up to this study, a successful effort was undertaken to agree on a single molecular typing approach to scale up the typing capacity, and improve quality assessment, by introducing proficiency testing for SRLs that intended to participate in advanced S. aureus surveillance. This effort provided the basis for the execution of a mutually agreed protocol using a standardized sampling frame and a quality-controlled genetic typing approach [12],[15], based on the sequencing of the variable region of the S. aureus spa gene (spa typing) [11]. Multilocus sequence typing (MLST) was also carried out on many strains allowing most of the prevalent spa types to be related to MLST sequence types. However, given the scope and ambition of this investigation, it is not surprising that the study still suffered from several operational shortcomings. In order to keep the amount of work manageable for the participating SRLs, it was decided to include about 20 laboratories that were demographically and geographically representative of each of the participating countries. This number is arbitrary and cannot equally represent small and large countries alike. Thus the precision of spatial scan statistics is reduced in areas where the density of laboratories is low. Laboratory enrolment based on population size would provide a more appropriate sampling strategy but would also impose a proportional and frequently unacceptable amount of work on SRLs in large countries if the sample size from small countries should remain meaningful. Even medium-sized countries such as Bulgaria, Finland, Greece, and Romania were unable to enrol the intended number of laboratories mainly for technical and logistic reasons. Naturally, the number of laboratories and isolates varied between countries and geo-demographic representation could be improved in future investigations. The original intention was to collect equal numbers of successive MSSA and MRSA in all laboratories during the 6-mo sampling interval; this proved to be unrealistic, especially in countries where MRSA levels are low. As a result, Norway and Iceland could not provide any MRSA, whereas Sweden, Denmark, and the Netherlands each provided fewer than ten isolates. Cyprus and Malta had only one participating laboratory but since both provide the microbiological service for the whole of the respective island population (for Cyprus, only the Republic of Cyprus), they were entitled to submit up to 20 isolates. Nevertheless, even taking these potential problems into account, the simultaneous collection of 2,890 isolates from patients with invasive S. aureus infection treated in 450 hospitals during a 6-mo study interval is unparalleled and remains unmatched by previous investigations, which have drawn their conclusions from convenience samples of predominantly MRSA collected by laboratories for different clinical or biological reasons. The current collection includes one-third MRSA and thus over-represents the natural population of MRSA causing invasive disease, which was 22% in the European Union in 2007 [24]. All isolates were collected from laboratories and hospitals participating in EARSS for which estimates of the overall catchment population are known. Thus, the present sample of hospitals catered for approximately 22 million people, totalling 4.4% of the citizens living in the European Union. Despite the above limitations, the sample provides a realistic insight into the epidemiology of S. aureus currently causing invasive infection in Europe. The age distribution and all-cause mortality was consistent with the expected range [29],[30]. High frequencies of invasive infections were ascertained in the very young (infants and under 5-y-olds) and older age groups with males clearly more at risk than females. Patients with MRSA were older than patients with MSSA and were 2.4 times more likely to have their infection attributed to hospital acquisition (p<0.0001). Invasive MRSA carried a higher all-cause mortality after 14 d. This finding is most likely confounded by a host of variables that distinguish MRSA patients from MSSA patients in fundamental ways. All-cause mortality was independent of spa type, indicating that this study did not identify any single spa type that stands out with respect to hyper-virulence or other factors that would increase the risk of fatal outcome after 14 d. While a laboratory-based cross-sectional study is limited in its ability to control for many of the crucial confounders such as comorbidity and disease severity scores and thus may not detect subtle differences in virulence properties between different clonal lineages at the patient level, this sample provides an unbiased estimate of the frequencies of specific spa types that have previously been reported to cause outbreaks and have attracted considerable public health attention. Few MRSA isolates carried the PVL-toxin genes and this could be an indication that many CO-MRSA were originally hospital-acquired. Only six of the PVL-positive CO-MRSA isolates, which made up 0.5% of the overall sample, had spa types consistent with the CA-MRSA clones most frequently reported in Europe (t008/ST8/SCCmecIV and t044/ST80/SCCmecIV) [31]. These values when compared to the overall numbers of MRSA are small but still warrant attention since PVL-positive CA-MRSA are more commonly associated with skin and soft tissue infections and are rarely found in blood stream infections [31] from which the majority of the sample isolates were drawn. Livestock-associated spa types belonging to MLST sequence type ST398 [32] made up 0.4% of the overall sample. However, none were methicillin-resistant indicating a low rate of human systemic infections induced by MRSA variants of this clone despite the increasing interest and concern about such isolates by health authorities. With the decision to utilize spa typing as a common platform to address the geographic abundance of S. aureus clones, various potential problems that might affect observations need to be taken into account. First, a single sequence of approximately 280 base-pairs under potential immune selection may be a relatively weak indicator for the genetic background of a genome that is approximately 10,000-times larger, even in a species such as S. aureus that is evolving in a predominantly clonal manner. Furthermore, convergent evolution could occur as a result of the high mutability of the repeat region of the spa gene used in spa typing [26],[33]. Finally, spa typing on its own it may not be sufficiently discriminatory to distinguish between MRSA isolates given that only five spa types accounted for almost half (48.1%) of all 967 MRSA isolates examined in this survey. The regional clusters found in this study provide a good indication that homoplasy is not a major problem that hinders the recognition of clonal dissemination on a geographic scale. Moreover, when comparing agr type, SCCmec type, toxin gene, and antibiotic susceptibility profiles within and between different spa types, a significant within-spa-type consistency and between-spa-type discordance supports the notion that in most cases spa typing provides a convenient and valid marker for the major clones and clonal lineages [34]. In order to explore the geographic distribution of different spa types, and the genetic and phenotypic detail of isolates in the European sample, the reader is encouraged to explore the purpose-built interactive Web application, SRL-Maps, at http://www.spatialepidemiology.net/srl-maps/. This application contains the public domain data made available to the scientific and public health community by all SRLs that participated in the study. It also illustrates the potential of such a communication platform. The underlying database is fully searchable and the map-based application is a template for the future addition of further epidemiological and biological information. We believe that this approach to spatial epidemiology will become a rich resource for future enquiry into the population dynamics of infectious agents and their evolution. In conclusion we provide evidence that the major MRSA clones in Europe occur predominantly in geographical clusters. This also indicates that MRSA, rather than spreading freely in the community, diffuses through regional health care networks. This important finding suggests that control efforts aimed at interrupting the spread within and between health care institutions may not only be feasible but ultimately successful and should therefore be strongly encouraged. We also showed that an international surveillance network sharing decentralised typing results on a Web-based platform can provide crucial information for clinicians, diagnostic microbiologists, and infection control teams on the dynamics of S. aureus spread, and especially the spread of MRSA isolates, to provide early warning of emerging strains, cross-border spread, and importation by travel. Supporting Information Text S1 Affiliations and contact information of the Staphylococcus aureus Reference Laboratory Working Group members. (0.16 MB DOC) Click here for additional data file. Text S2 Author contributions of the Staphylococcus aureus Reference Laboratory Working Group members. (0.13 MB DOC) Click here for additional data file.