- Record: found
- Abstract: found
- Article: found
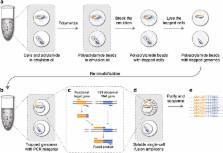
Massively parallel sequencing of single cells by epicPCR links functional genes with phylogenetic markers
Read this article at
Abstract
Many microbial communities are characterized by high genetic diversity. 16S ribosomal RNA sequencing can determine community members, and metagenomics can determine the functional diversity, but resolving the functional role of individual cells in high throughput remains an unsolved challenge. Here, we describe epicPCR (Emulsion, Paired Isolation and Concatenation PCR), a new technique that links functional genes and phylogenetic markers in uncultured single cells, providing a throughput of hundreds of thousands of cells with costs comparable to one genomic library preparation. We demonstrate the utility of our technique in a natural environment by profiling a sulfate-reducing community in a freshwater lake, revealing both known sulfate reducers and discovering new putative sulfate reducers. Our method is adaptable to any conserved genetic trait and translates genetic associations from diverse microbial samples into a sequencing library that answers targeted ecological questions. Potential applications include identifying functional community members, tracing horizontal gene transfer networks and mapping ecological interactions between microbial cells.
Related collections
Most cited references23
- Record: found
- Abstract: found
- Article: not found
The ecology and biotechnology of sulphate-reducing bacteria.
- Record: found
- Abstract: found
- Article: not found
Microfluidic digital PCR enables multigene analysis of individual environmental bacteria.

- Record: found
- Abstract: found
- Article: found