- Record: found
- Abstract: found
- Article: not found
Golden Gate Shuffling: A One-Pot DNA Shuffling Method Based on Type IIs Restriction Enzymes
Read this article at
Abstract
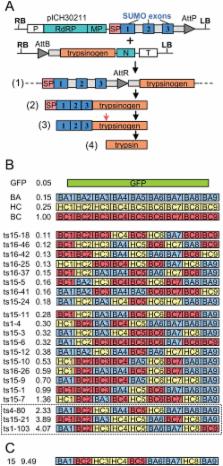
We have developed a protocol to assemble in one step and one tube at least nine separate DNA fragments together into an acceptor vector, with 90% of recombinant clones obtained containing the desired construct. This protocol is based on the use of type IIs restriction enzymes and is performed by simply subjecting a mix of 10 undigested input plasmids (nine insert plasmids and the acceptor vector) to a restriction-ligation and transforming the resulting mix in competent cells. The efficiency of this protocol allows generating libraries of recombinant genes by combining in one reaction several fragment sets prepared from different parental templates. As an example, we have applied this strategy for shuffling of trypsinogen from three parental templates (bovine cationic trypsinogen, bovine anionic trypsinogen and human cationic trypsinogen) each divided in 9 separate modules. We show that one round of shuffling using the 27 trypsinogen entry plasmids can easily produce the 19,683 different possible combinations in one single restriction-ligation and that expression screening of a subset of the library allows identification of variants that can lead to higher expression levels of trypsin activity. This protocol, that we call ‘Golden Gate shuffling’, is robust, simple and efficient, can be performed with templates that have no homology, and can be combined with other shuffling protocols in order to introduce any variation in any part of a given gene.
Related collections
Most cited references41
- Record: found
- Abstract: found
- Article: not found
A One Pot, One Step, Precision Cloning Method with High Throughput Capability
- Record: found
- Abstract: found
- Article: not found
Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC.
- Record: found
- Abstract: found
- Article: not found
