- Record: found
- Abstract: found
- Article: found
Phylogenomic resolution of Imparidentia (Mollusca: Bivalvia) diversification through mitochondrial genomes
Abstract
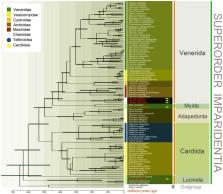
Despite significant advances in the phylogenomics of bivalves over the past decade, the higher-level phylogeny of Imparidentia (a superorder of Heterodonta) remains elusive. Here, a total of five new mitochondrial sequences ( Chama asperella, Chama limbula, Chama dunkeri, Barnea manilensis and Ctena divergens) was added to provide resolution in nodes that required additional study . Although the monophyly of Lucinida remains less clear, the results revealed the overall backbone of the Imparidentia tree and the monophyly of Imparidentia. Likewise, most relationships among the five major Imparidentia lineages—Lucinida, Cardiida, Adapedonta, Myida and Venerida—were addressed with a well-supported topology. Basal relationships of Imparidentia recovered Lucinidae as the sister group to all remaining imparidentian taxa. Thyasiridae is a sister group to other imparidentian bivalves (except Lucinidae species) which is split into Cardiida, Adapedonta and the divergent clade of Neoheterodontei. Neoheterodontei was comprised of Venerida and Myida, the former of which now also contains Chamidae as the sister group to all the remaining venerid taxa. Moreover, molecular divergence times were inferred by calibrating nine nodes in the Imparidentia tree of life by extinct taxa. The origin of these major clades ranged from Ordovician to Permian with the diversification through the Palaeozoic to Mesozoic. Overall, the results obtained in this study demonstrate a better-resolved Imparidentia phylogeny based on mitochondrial genomes.
Related collections
Most cited references64
- Record: found
- Abstract: found
- Article: not found
MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets.

- Record: found
- Abstract: found
- Article: found
IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies

- Record: found
- Abstract: found
- Article: found