- Record: found
- Abstract: found
- Article: found
CircPrimer 2.0: a software for annotating circRNAs and predicting translation potential of circRNAs

Read this article at
Abstract
Background
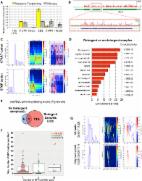
Some circular RNAs (circRNAs) can be translated into functional peptides by small open reading frames (ORFs) in a cap-independent manner. Internal ribosomal entry site (IRES) and N 6-methyladenosine (m 6A) were reported to drive translation of circRNAs. Experimental methods confirming the presence of IRES and m 6A site are time consuming and labor intensive. Lacking computational tools to predict ORFs, IRESs and m 6A sites for circRNAs makes it harder.
Results
In this report, we present circPrimer 2.0, a Java based software for annotating circRNAs and predicting ORFs, IRESs, and m6A sites of circRNAs. circPrimer 2.0 has a graphical and a command-line interface that enables the tool to be embed into an analysis pipeline.
Conclusions
circprimer 2.0 is an easy-to-use software for annotating circRNAs and predicting translation potential of circRNAs, and freely available at www.bio-inf.cn.
Related collections
Most cited references28

- Record: found
- Abstract: found
- Article: not found
Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis

- Record: found
- Abstract: found
- Article: found
Extensive translation of circular RNAs driven by N6-methyladenosine