- Record: found
- Abstract: found
- Article: not found
Current trends in COVID-19 diagnosis and its new variants in physiological fluids: Surface antigens, antibodies, nucleic acids, and RNA sequencing

Read this article at
Abstract
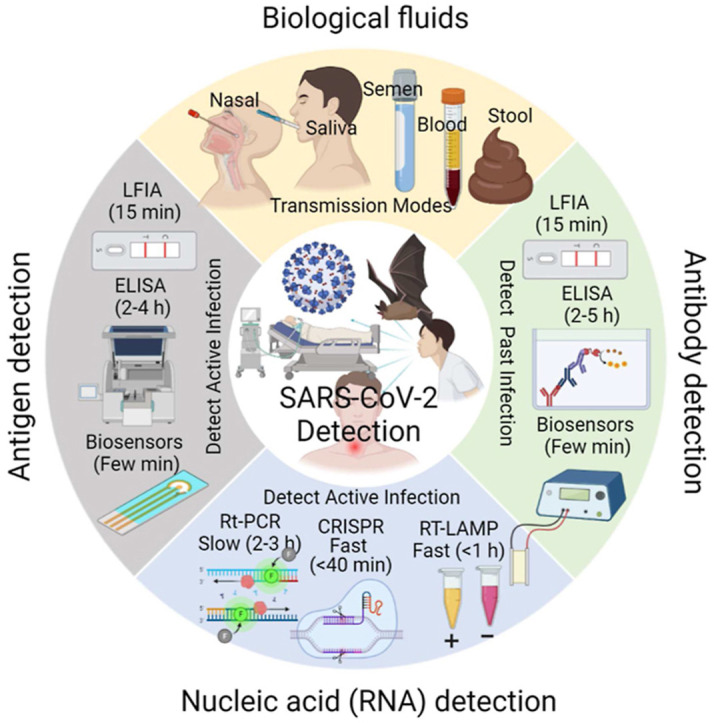
Rapid, highly sensitive, and accurate virus circulation monitoring techniques are critical to limit the spread of the virus and reduce the social and economic burden. Therefore, point-of-use diagnostic devices have played a critical role in addressing the outbreak of COVID-19 (SARS-CoV-2) viruses. This review provides a comprehensive overview of the current techniques developed for the detection of SARS-CoV-2 in various body fluids (e.g., blood, urine, feces, saliva, tears, and semen) and considers the mutations (i.e., Alpha, Beta, Gamma, Delta, Omicron). We classify and comprehensively discuss the detection methods depending on the biomarker measured (i.e., surface antigen, antibody, and nucleic acid) and the measurement techniques such as lateral flow immunoassay (LFIA), enzyme-linked immunosorbent assay (ELISA), reverse transcriptase-polymerase chain reaction (RT-PCR), reverse transcription loop-mediated isothermal amplification (RT-LAMP), microarray analysis, clustered regularly interspaced short palindromic repeats (CRISPR) and biosensors. Finally, we addressed the challenges of rapidly identifying emerging variants, detecting the virus in the early stages of infection, the detection sensitivity, selectivity, and specificity, and commented on how these challenges can be overcome in the future.
Graphical abstract
Related collections
Most cited references161
- Record: found
- Abstract: found
- Article: not found
A Novel Coronavirus from Patients with Pneumonia in China, 2019
- Record: found
- Abstract: found
- Article: found
Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus–Infected Pneumonia in Wuhan, China
- Record: found
- Abstract: found
- Article: not found