- Record: found
- Abstract: found
- Article: found
Proteomic profiling of glucocorticoid-exposed myogenic cells: Time series assessment of protein translocation and transcription of inactive mRNAs
Read this article at
Abstract
Background
Prednisone, one of the most highly prescribed drugs, has well characterized effects on gene transcription mediated by the glucocorticoid receptor. These effects are typically occurring on the scale of hours. Prednisone also has a number of non-transcriptional effects (occurring on minutes scale) on protein signaling, yet these are less well studied. We sought to expand the understanding of acute effects of prednisone action on cell signaling using a combination of SILAC strategy and subcellular fractionations from C 2C 12 myotubes.
Results
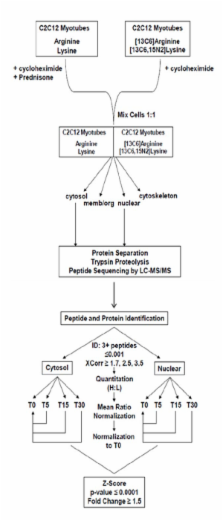
De novo translation of proteins was inhibited in both SILAC labeled and unlabeled C 2C 12 myotubes. Unlabeled cells were exposed to prednisone while SILAC labeled cells remained untreated. After 0, 5, 15, and 30 minutes of prednisone exposure, labeled and unlabeled cells were mixed at 1:1 ratios and fractionated into cytosolic and nuclear fractions. A total of 534 proteins in the cytosol and 626 proteins in the nucleus were identified and quantitated, using 3 or more peptides per protein with peptide based probability ≤ 0.001. We identified significant increases (1.7- to 3.1- fold) in cytoplasmic abundance of 11 ribosomal proteins within 5 minutes of exposure, all of which returned to baseline by 30 min. We hypothesized that these drug-induced acute changes in the subcellular localization of the cell's protein translational machinery could lead to altered translation of quiescent RNAs. To test this, de novo protein synthesis was assayed after 15 minutes of drug exposure. Quantitative fluorography identified 16 2D gel spots showing rapid changes in translation; five of these were identified by MS/MS (pyruvate kinase, annexin A6 isoform A and isoform B, nasopharyngeal epithelium specific protein 1, and isoform 2 of Replication factor C subunit 1), and all showed the 5' terminal oligopyrimidine motifs associated with mRNA sequestration to and from inactive mRNA pools.
Conclusion
We describe novel approaches of subcellular proteomic profiling and assessment of acute changes on a minute-based time scale. These data expand the current knowledge of acute, non-transcriptional activities of glucocorticoids, including changes in protein subcellular localization, altered translation of quiescent RNA pools, and PKC-mediated cytoskeleton remodeling.
Related collections
Most cited references46
- Record: found
- Abstract: found
- Article: not found
Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin.
- Record: found
- Abstract: found
- Article: not found