- Record: found
- Abstract: found
- Article: found
Can Biofilm Be Reversed Through Quorum Sensing in Pseudomonas aeruginosa?
Read this article at
Abstract
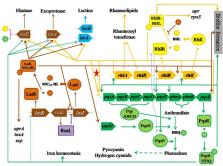
Pseudomonas aeruginosa is a Gram-negative bacterium causing diseases in plants, animals, and humans, and its drug resistance is a major concern in medical care. Biofilms play an important role in P. aeruginosa drug resistance. Three factors are most important to induce biofilm: quorum sensing (QS), bis-(3′-5′)-cyclic diguanosine monophosphate (c-di-GMP), and small RNAs (sRNAs). P. aeruginosa has its own specific QS system (PQS) besides two common QS systems, LasI–LasR and RhlI–RhlR, in bacteria. PQS is interesting not only because there is a negative regulation from RhlR to pqsR but also because the null mutation in PQS leads to a reduced biofilm formation. Furthermore, P. aeruginosa dispersed cells have physiological features that are distinct between the planktonic cells and biofilm cells. In response to a low concentration of c-di-GMP, P. aeruginosa cells can disperse from the biofilms to become planktonic cells. These raise an interesting hypothesis of whether biofilm can be reversed through the QS mechanism in P. aeruginosa. Although a single factor is certainly not sufficient to prevent the biofilm formation, it necessarily explores such possibility. In this hypothesis, the literature is analyzed to determine the negative regulation pathways, and then the transcriptomic data are analyzed to determine whether this hypothesis is workable or not. Unexpectedly, the transcriptomic data reveal a negative regulation between lasI and psqR. Also, the individual cases from transcriptomic data demonstrate the negative regulations of PQS with laslI, laslR, rhlI, and rhlR under different experiments. Based on our analyses, possible strategies to reverse biofilm formation are proposed and their clinic implications are addressed.
Related collections
Most cited references86
- Record: found
- Abstract: found
- Article: not found
The hierarchy quorum sensing network in Pseudomonas aeruginosa
- Record: found
- Abstract: not found
- Article: not found
Quorum sensing in bacteria: the LuxR-LuxI family of cell density-responsive transcriptional regulators.
- Record: found
- Abstract: found
- Article: not found