- Record: found
- Abstract: found
- Article: not found
Influence of age, gender, previous SARS-CoV-2 infection, and pre-existing diseases in antibody response after COVID-19 vaccination: A review
review-article

Read this article at
There is no author summary for this article yet. Authors can add summaries to their articles on ScienceOpen to make them more accessible to a non-specialist audience.
Abstract
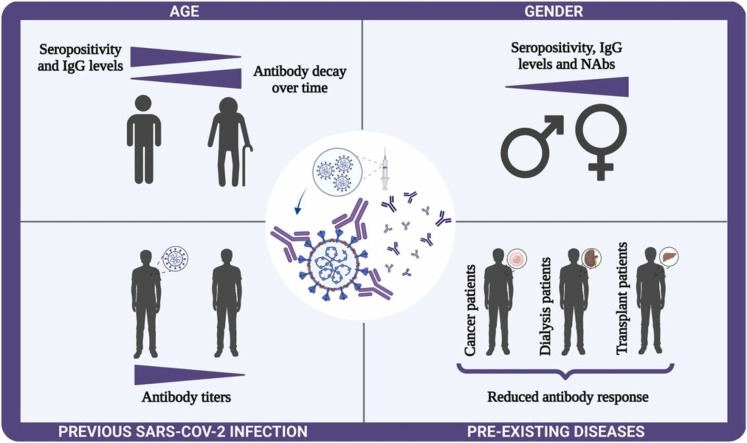
Vaccines induce specific long-term immunological memory against pathogens, preventing the worsening of diseases. The COVID-19 health emergency has caused more than 6 million deaths and started a race for vaccine development. Antibody response to COVID-19 vaccines has been investigated primarily in healthcare workers. The heterogeneity of immune responses and the behavior of this response in particular groups were still very little explored. In this review, we discuss whether antibody responses after vaccination are influenced by age, gender, previous SARS-CoV-2 infection, or pre-existing diseases.
Graphical Abstract
Related collections
Most cited references75
- Record: found
- Abstract: found
- Article: not found
Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding
Roujian Lu, Xiang Zhao, Juan Li … (2021)
- Record: found
- Abstract: found
- Article: not found
The Architecture of SARS-CoV-2 Transcriptome
Dongwan Kim, Joo-Yeon Lee, Jeong-Sun Yang … (2020)