- Record: found
- Abstract: found
- Article: found
NK Cells Stimulate Recruitment of cDC1 into the Tumor Microenvironment Promoting Cancer Immune Control

Read this article at
Summary
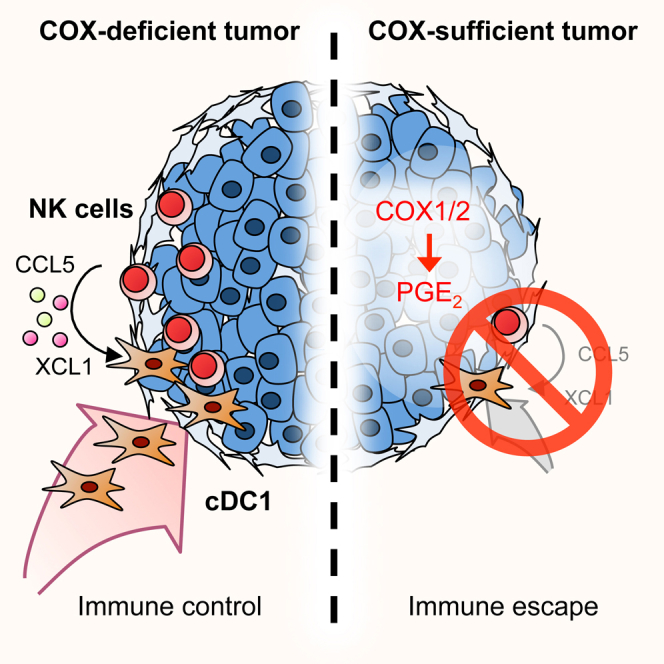
Conventional type 1 dendritic cells (cDC1) are critical for antitumor immunity, and their abundance within tumors is associated with immune-mediated rejection and the success of immunotherapy. Here, we show that cDC1 accumulation in mouse tumors often depends on natural killer (NK) cells that produce the cDC1 chemoattractants CCL5 and XCL1. Similarly, in human cancers, intratumoral CCL5, XCL1, and XCL2 transcripts closely correlate with gene signatures of both NK cells and cDC1 and are associated with increased overall patient survival. Notably, tumor production of prostaglandin E2 (PGE 2) leads to evasion of the NK cell-cDC1 axis in part by impairing NK cell viability and chemokine production, as well as by causing downregulation of chemokine receptor expression in cDC1. Our findings reveal a cellular and molecular checkpoint for intratumoral cDC1 recruitment that is targeted by tumor-derived PGE 2 for immune evasion and that could be exploited for cancer therapy.
Graphical Abstract
Highlights
Abstract
Natural killer cells recruit dendritic cells to the tumor microenvironment, and disruption of this process results in cancer immune evasion.
Related collections
Most cited references30
- Record: found
- Abstract: found
- Article: not found
Batf3 deficiency reveals a critical role for CD8alpha+ dendritic cells in cytotoxic T cell immunity.
- Record: found
- Abstract: found
- Article: not found
Innate Immune Landscape in Early Lung Adenocarcinoma by Paired Single-Cell Analyses
- Record: found
- Abstract: found
- Article: not found