- Record: found
- Abstract: found
- Article: found
Editorial: Engineering Corynebacterium glutamicum Chassis for Synthetic Biology, Biomanufacturing, and Bioremediation
editorial
Yu Wang
1 ,
Ping Zheng
1
,
*
,
,
Zhenghong Xu
2 ,
Akihiko Kondo
3 ,
Christoph Wittmann
4 ,
Volker F. Wendisch
5
,
*
,
08 June 2022
Read this article at
There is no author summary for this article yet. Authors can add summaries to their articles on ScienceOpen to make them more accessible to a non-specialist audience.
Abstract
The Gram-positive soil bacterium Corynebacterium glutamicum was discovered about 60 years
ago as an l-glutamate producer and has become a leading workhorse in industrial biotechnology.
It is now used for the industrial production of over 6 million tons of amino acids
(such as l-lysine and l-glutamate) per year and shows great potential for producing
many more compounds ranging from alcohols and organic acids to plant secondary metabolites
(Wolf et al., 2021). Several characteristics of C. glutamicum make it particularly
interesting for industrial biotechnology, such as the GRAS (generally regarded as
safe) status of its products, fast growth with relatively few nutrient requirements,
and capability of utilizing sugars, sugar alcohols, organic acids, and aromatic compounds.
With the development of CRISPR-based genome editing methods and synthetic and systems
biology tools, the ability to understand and engineer the metabolism and regulation
of C. glutamicum has been extensively enhanced (Liu et al., 2022). However, over 40%
of genes of the type strain ATCC 13032 have not been experimentally characterized
for their biological functions and the metabolic and regulatory mechanisms underlying
the superior industrial performance of C. glutamicum have not been fully deciphered.
Research on metabolic modeling, chassis engineering, multi-omics understanding, genome
mining, etc., is expected to further realize the potential of this bacterium in biomanufacturing
of chemicals and proteins and bioremediation of pollutants.
A Research Topic of articles including 18 original research articles and 1 mini review
article specialized in C. glutamicum research from the leading groups in this field
is presented. Each article provides a state-of-the-art view of the metabolic engineering
efforts, systems biology analyses, and/or technical advances. The Research Topic focuses
on the development of enabling technologies, mining of functional components, and
engineering of C. glutamicum as microbial cell factories for bioconversion of renewable
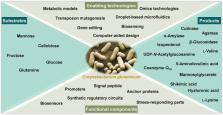
feedstocks to useful chemicals and proteins (Figure 1).
FIGURE 1
A general categorization of the keywords of the articles collected in this Research
Topic.
It is great to see that a significant part of the Research Topic presents modern enabling
technologies and their practical application in C. glutamicum. In the design-build-test-learn
biological engineering cycle, genome-scale metabolic models (GEMs) are considered
fundamental tools. Feierabend et al. presented an updated and unified GEM of C. glutamicum
ATCC 13032 (iCGB21FR) with high quality regarding comprehensiveness and data standards.
The in silico analysis may provide numerous designs for strain engineering. To bring
these designs to life, computer-aided design tools and robotic system-assisted genome
editing technologies will certainly help. Yang et al. developed a free online tool
called GEDpm-cg for the design of genomic point mutations in C. glutamicum. Primers
required for tool plasmid assembly and sequencing verification were quickly designed
and provided, which would be useful for large-scale mutation analysis. As summarized
by Wang et al., versatile genome editing tools including those based on the CRIPSR
systems have been developed for C. glutamicum. Tailored design tools for not only
point mutation but also large-scale genome engineering are still needed. In the test
and learn parts, many high-throughput and systematic technologies including genome-scale
transposon mutagenesis (Linder et al.), droplet-based microfluidics (Balasubramanian
et al.), biosensing (Bakkes et al.), and multi-omics analyses (Kappelmann et al. and
Banerjee et al.) have been used. Kappelmann et al. performed a comprehensive analysis
of single or double deletion mutants in the anaplerosis of C. glutamicum under defined
glucose conditions. Valuable information regarding the genotype-phenotype relationships
in these mutants was unraveled by combining untargeted proteomics, quantitative metabolomics,
and whole-genome sequencing. Banerjee et al. conducted genome and RNA sequencing of
an engineered isopentenol-producing C. glutamicum strain under industrially relevant
conditions including scale transition and ionic liquid stress. This omics information
clarified the cell response of a C. glutamicum strain engineered to produce isopentenol.
The advances of enabling technologies considerably accelerate the running of the design-build-test-learn
cycle and promote the engineering of C. glutamicum chassis, development of new components
and circuits, and construction of microbial cell factories. To engineer a chassis
that can efficiently metabolize fructose and channel the carbon flux to the oxidative
pentose phosphate pathway for NADHP generation, Krahn et al. engineered and evolved
fructose-utilizing mutants. Crucial mutations in the glucose phosphotransferase system
enzymes were identified to explain the altered fructose uptake. A C. glutamicum chassis
named CR101 with the removal of prophages and all insertion sequence (IS) elements
was constructed by Linder et al. This chassis CR101 shows growth characteristics identical
to the wild-type and increased transformability and could serve as an optimal host
for basic research and biotechnology including genome-scale transposon mutagenesis.
Several articles collected in this Research Topic describe new functional components
for engineering of C. glutamicum. To provide more available anchoring motifs for the
display of recombinant proteins on the surfaces of C. glutamicum cells, Lin et al.
predicted and screened 14 potential anchor proteins and identified 3 new anchoring
proteins that performed better than the commonly used ones. Inducible gene expression
systems are always important for reprogramming the metabolism and regulation of microorganisms.
Three sets of gene regulation systems induced by myo-inositol, hyperosmotic stress,
and phenolic compounds such as ferulic acid were developed and applied for dynamic
control of gene expression. Lu et al. constructed a myo-inositol-inducible expression
vector pMI-4 using the iolR
q
cassette and the myo-inositol-inducible promoter P
iolT1
, which was used for gene overexpression for 5-aminolevulinic acid production. Huang
et al. characterized the promoters controlled by the two-component signal transduction
system MtrA/MtrB responding to hyperosmotic stress. The promoter of NCgl1418 that
exhibited a high inducibility was further engineered and used to develop a CRISRPi
system induced by hyperosmotic stress. Siebert et al. utilized the VanR/P
vanABK
* regulatory system responding to phenolic compounds ferulic acid, vanillin, and vanillic
acid to develop a timed off-switch for dynamic control of gene expression. With the
depletion of exogenous phenolic compounds, genes controlled by P
vanABK
* were efficiently turned off, allowing the control of gene expression in C. glutamicum
in a timed manner.
Finally, a substantial part of this article Research Topic reports bioproduction of
chemicals and proteins by C. glutamicum. Sato et al. engineered a C. glutamicum strain
with β-glucosidase secreting ability for production of shikimate from both glucose
and cellobiose. The shikimate pathway is a common route for the biosynthesis of a
range of aromatic compounds, which also provides precursors for the biosynthesis of
coenzyme Q10, an electron carrier in aerobic respiration and an antioxidant in medical
treatment. Burgardt et al. metabolically engineered C. glutamicum for de novo biosynthesis
of coenzyme Q10, which to the best of our knowledge is the first report of coenzyme
Q10 production in a non-ubiquinone-containing bacterium. Uridine diphosphate-N-acetylglucosamine
(UDP-GlcNAc) is an acetylated amino sugar nucleotide that can serve as a sugar donor
for synthesis of many pharmaceutically relevant oligosaccharides, polysaccharides,
and glycoproteins. Gauttam et al. constructed a series of recombinant C. glutamicum
strains by engineering the UDP-GlcNAc biosynthetic pathway and the highest level of
microbial production of UDP-GlcNAc was obtained. Different from most metabolic engineering
strategies directly targeting the biosynthetic pathway, Du et al. applied indirect
metabolic engineering strategies targeting the substrate uptake, membrane composition,
oxygen transfer, and nitrogen metabolism for enhanced biosynthesis of hyaluronic acid.
Metabolic engineering approaches have also been combined with process engineering
approaches by Schwentner et al. to develop a C. glutamicum strain for the production
of compatible solute mannosylglycerate and an easy product separation process to extract
mannosylglycerate from cytosol by cold water shock.
In addition to bioproduction of chemicals, secretory production of recombinant proteins
using C. glutamicum also attracts great attention. Lee et al. found that mutations
to increase the iron and carbon consumption were responsible for the enhanced production
of recombinant protein in an evolved fast-growing C. glutamicum strain. Balasubramanian
et al. screened C. glutamicum mutants with enhanced enzyme secretion capacity by using
a droplet-based microfluidic high-throughput screening method and analyzed the single
nucleotide variants in these mutants. Bakkes et al. used a fluorescence-based biosensor
for Sec-dependent protein secretion to evolve the Sec signal peptides from Bacillus
subtilis and optimize the secretion of heterologous enzyme cutinase. These findings
are expected to promote the application of C. glutamicum in protein production.
Overall, C. glutamicum, that was initially used as an l-glutamate producer, is now
becoming a versatile chassis for bioproduction of various amino acids, chemicals,
and proteins from renewable feedstocks. The importance of C. glutamicum in industrial
biotechnology drives the development of advanced synthetic and systems biology technologies
such as in silico metabolic modeling, multiplex gene editing, multi-omics analyses,
and high-throughput functional genomics methods. It is believed that the series of
articles collected in this Research Topic provide contemporary and valuable information
that can help to fully unlock the potential of C. glutamicum in industrial biotechnology.
Related collections
Most cited references2

- Record: found
- Abstract: found
- Article: found
Advances in metabolic engineering of Corynebacterium glutamicum to produce high-value active ingredients for food, feed, human health, and well-being
Sabrina Wolf, Judith Becker, Yota Tsuge … (2021)

- Record: found
- Abstract: found
- Article: found