- Record: found
- Abstract: found
- Article: not found
Structural biology of the Mre11:Nbs1 complex structure yields insights into ataxia–telangiectasia–like disease mutations and DNA damage signaling
Read this article at
Summary
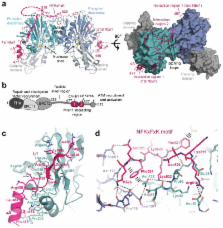
The Mre11–Rad50–Nbs1 (MRN) complex tethers, processes and signals DNA double strand breaks, promoting genomic stability. To understand the functional architecture of MRN, we determined the crystal structures of the Schizosaccharomyces pombe Mre11 dimeric catalytic domain alone and in complex with a fragment of Nbs1. Two Nbs1 subunits stretch around the outside of Mre11’s nuclease domains, with one subunit additionally bridging and locking the Mre11 dimer via a highly conserved asymmetrical binding motif. Our results reveal that Mre11 forms a flexible dimer and suggest that Nbs1 is not only a checkpoint adaptor, but also functionally impacts on Mre11-Rad50. Clinical mutations in Mre11 are located along the Nbs1 interaction sites and weaken the Mre11–Nbs1 interaction. However, they differentially affect DNA repair and telomere maintenance in Saccharomyces cerevisiae, potentially providing insight into their different human disease pathologies.
Related collections
Most cited references56
- Record: found
- Abstract: found
- Article: not found
The DNA damage response: ten years after.
- Record: found
- Abstract: found
- Article: not found
Human CtIP promotes DNA end resection.
- Record: found
- Abstract: found
- Article: not found
