- Record: found
- Abstract: found
- Article: found
The integrative future of taxonomy
Read this article at
Abstract
Background
Taxonomy is the biological discipline that identifies, describes, classifies and names extant and extinct species and other taxa. Nowadays, species taxonomy is confronted with the challenge to fully incorporate new theory, methods and data from disciplines that study the origin, limits and evolution of species.
Results
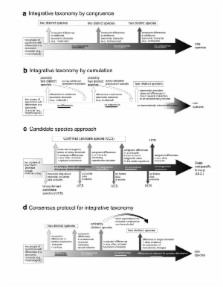
Integrative taxonomy has been proposed as a framework to bring together these conceptual and methodological developments. Here we review perspectives for an integrative taxonomy that directly bear on what species are, how they can be discovered, and how much diversity is on Earth.
Conclusions
We conclude that taxonomy needs to be pluralistic to improve species discovery and description, and to develop novel protocols to produce the much-needed inventory of life in a reasonable time. To cope with the large number of candidate species revealed by molecular studies of eukaryotes, we propose a classification scheme for those units that will facilitate the subsequent assembly of data sets for the formal description of new species under the Linnaean system, and will ultimately integrate the activities of taxonomists and molecular biologists.
Related collections
Most cited references73
- Record: found
- Abstract: found
- Article: not found
Cryptic species as a window on diversity and conservation.
- Record: found
- Abstract: found
- Article: not found
Linkage disequilibrium--understanding the evolutionary past and mapping the medical future.
- Record: found
- Abstract: found
- Article: not found