- Record: found
- Abstract: found
- Article: found
The lncRNA Toolkit: Databases and In Silico Tools for lncRNA Analysis
Read this article at
Abstract
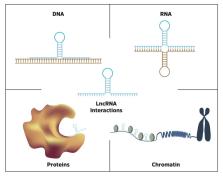
Long non-coding RNAs (lncRNAs) are a rapidly expanding field of research, with many new transcripts identified each year. However, only a small subset of lncRNAs has been characterized functionally thus far. To aid investigating the mechanisms of action by which new lncRNAs act, bioinformatic tools and databases are invaluable. Here, we review a selection of computational tools and databases for the in silico analysis of lncRNAs, including tissue-specific expression, protein coding potential, subcellular localization, structural conformation, and interaction partners. The assembled lncRNA toolkit is aimed primarily at experimental researchers as a useful starting point to guide wet-lab experiments, mainly containing multi-functional, user-friendly interfaces. With more and more new lncRNA analysis tools available, it will be essential to provide continuous updates and maintain the availability of key software in the future.
Related collections
Most cited references172
- Record: found
- Abstract: found
- Article: not found
Basic local alignment search tool.
- Record: found
- Abstract: found
- Article: not found
Proteomics. Tissue-based map of the human proteome.

- Record: found
- Abstract: found
- Article: found