- Record: found
- Abstract: found
- Article: found
The Microbiota Dynamics of Alfalfa Silage During Ensiling and After Air Exposure, and the Metabolomics After Air Exposure Are Affected by Lactobacillus casei and Cellulase Addition
Read this article at
Abstract
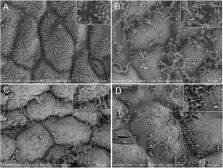
Both inoculants treatment and enzyme treatment promote the reproduction of lactic acid bacteria (LAB) to produce enough lactic acid to lower pH in silage. The present study investigated the microbial community and metabolome in cellulase, Lactobacillus casei, and air treated alfalfa silage. Chopped and wilted alfalfa (first cutting, 29% dry matter) was ensiled without (CON) or with L. casei (1 × 10 6 cfu g –1 fresh matter) (LC) or cellulase (20,000IU, 0.5% of fresh matter) (CE) for 56 days, then exposed to air for 3 days (PO). Greater ensiling quality was observed in LC and CE, which had lower pH and higher lactic acid content than CON at 56 days of ensiling and 3 days post-oxygen exposure. Air exposure was associated with decreased lactic acid concentrations and increased yeast and mold counts in all silages. SEM showed that the structure of leaf epicuticular wax crystals were intact in fresh alfalfa, totally decomposed in CON silage, and partly preserved in CE and LC silage. Gas chromatography mass spectrometry revealed that 196 metabolites and 95 differential concentration were present in the 3 days air exposure samples. Most of these metabolites, mainly organic acids, polyols, ketones, aldehydes, are capable of antimicrobial activity. The bacterial communities were obviously different among groups and Lactobacillus developed to a dominant status in all silages. Lactobacillus became dominant in bacterial communities of LC and CE silages from days 7 to 56, and their relative abundances reached 94.17–83.93% at day 56, respectively. For CON silage, until day 56, Lactobacillus dominated the bacterial community with abundance of 75.10%. After 3 days of oxygen exposure, Lactobacillus and Enterococcus were predominant in CON, and Lactobacillus remained dominant in LC and CE silages. The results indicated that, compared to untreated silages, L. casei could be a priority inoculant for alfalfa silage to boost Lactobacillus abundance and improve fermentation quality. Our high-throughput sequencing and gas chromatography mass spectrometry results provide a deep insight into the bacterial community and metabolites in alfalfa silage.
Related collections
Most cited references64

- Record: found
- Abstract: found
- Article: found
Trimmomatic: a flexible trimmer for Illumina sequence data
- Record: found
- Abstract: not found
- Article: not found
QIIME allows analysis of high-throughput community sequencing data.
- Record: found
- Abstract: found
- Article: not found