- Record: found
- Abstract: found
- Article: found
Population‐level inferences from environmental DNA—Current status and future perspectives
Read this article at
Abstract
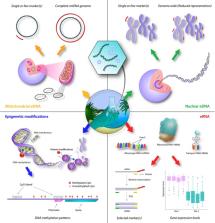
Environmental DNA (eDNA) extracted from water samples has recently shown potential as a valuable source of population genetic information for aquatic macroorganisms. This approach offers several potential advantages compared with conventional tissue‐based methods, including the fact that eDNA sampling is noninvasive and generally more cost‐efficient. Currently, eDNA approaches have been limited to single‐marker studies of mitochondrial DNA (mtDNA), and the relationship between eDNA haplotype composition and true haplotype composition still needs to be thoroughly verified. This will require testing of bioinformatic and statistical software to correct for erroneous sequences, as well as biases and random variation in relative sequence abundances. However, eDNA‐based population genetic methods have far‐reaching potential for both basic and applied research. In this paper, we present a brief overview of the achievements of eDNA‐based population genetics to date, and outline the prospects for future developments in the field, including the estimation of nuclear DNA (nuDNA) variation and epigenetic information. We discuss the challenges associated with eDNA samples as opposed to those of individual tissue samples and assess whether eDNA might offer additional types of information unobtainable with tissue samples. Lastly, we provide recommendations for determining whether an eDNA approach would be a useful and suitable choice in different research settings. We limit our discussion largely to contemporary aquatic systems, but the advantages, challenges, and perspectives can to a large degree be generalized to eDNA studies with a different spatial and temporal focus.
Related collections
Most cited references162
- Record: found
- Abstract: found
- Article: not found
Genome sequence of the nematode C. elegans: a platform for investigating biology.
- Record: found
- Abstract: found
- Article: not found
Molecular signatures of natural selection.
- Record: found
- Abstract: found
- Article: not found