- Record: found
- Abstract: found
- Article: found
H3K27 acetylation activated-long non-coding RNA CCAT1 affects cell proliferation and migration by regulating SPRY4 and HOXB13 expression in esophageal squamous cell carcinoma
Read this article at
Abstract
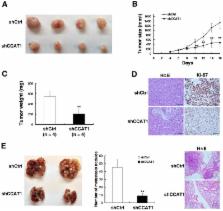
Recently, long non-coding RNAs (lncRNAs) have been shown to have important regulatory roles in human cancer biology. In our study, we found that lncRNA CCAT1, whose expression is significantly increased and is correlated with outcomes in Esophageal Squamous Cell Carcinoma (ESCC). Consecutive experiments confirmed that H3K27-acetylation could activate expression of colon cancer associated transcript-1 (CCAT1). Further experiments revealed that CCAT1 knockdown significantly repressed the proliferation and migration both in vitro and in vivo. RNA-seq analysis revealed that CCAT1 knockdown preferentially affected genes that are linked to cell proliferation, cell migration and cell adhesion. Mechanistic investigations found that CCAT1 could serve as a scaffold for two distinct epigenetic modification complexes (5΄ domain of CCAT1 binding Polycomb Repressive Complex 2 (PRC2) while 3΄ domain of CCAT1 binding SUV39H1) and modulate the histone methylation of promoter of SPRY4 (sprouty RTK signaling antagonist 4) in nucleus. In cytoplasm, CCAT1 regulates HOXB13 as a molecular decoy for miR-7, a microRNA that targets both CCAT1 and HOXB13, thus facilitating cell growth and migration. Together, our data demonstrated the important roles of CCAT1 in ESCC oncogenesis and might serve as targets for ESCC diagnosis and therapy.
Related collections
Most cited references28
- Record: found
- Abstract: found
- Article: not found
The transcriptional landscape of the mammalian genome.
- Record: found
- Abstract: found
- Article: not found
Long noncoding RNA as modular scaffold of histone modification complexes.
- Record: found
- Abstract: found
- Article: not found
