- Record: found
- Abstract: found
- Article: found
featureCounts: An efficient general-purpose program for assigning sequence reads to genomic features
Preprint
2013-05-14
Read this article at
There is no author summary for this article yet. Authors can add summaries to their articles on ScienceOpen to make them more accessible to a non-specialist audience.
Abstract
Next-generation sequencing technologies generate millions of short sequence reads,
which are usually aligned to a reference genome. In many applications, the key information
required for downstream analysis is the number of reads mapping to each genomic feature,
for example to each exon or each gene. The process of counting reads is called read
summarization. Read summarization is required for a great variety of genomic analyses
but has so far received relatively little attention in the literature. We present
featureCounts, a read summarization program suitable for counting reads generated
from either RNA or genomic DNA sequencing experiments. featureCounts implements highly
efficient chromosome hashing and feature blocking techniques. It is considerably faster
than existing methods (by an order of magnitude for gene-level summarization) and
requires far less computer memory. It works with either single or paired-end reads
and provides a wide range of options appropriate for different sequencing applications.
featureCounts is available under GNU General Public License as part of the Subread
(http://subread.sourceforge.net) or Rsubread (http://www.bioconductor.org) software
packages.
Related collections
Most cited references11

- Record: found
- Abstract: found
- Article: found
NCBI Reference Sequences (RefSeq): current status, new features and genome annotation policy
Kim D. Pruitt, Tatiana Tatusova, Garth R. Brown … (2011)

- Record: found
- Abstract: found
- Article: found
Comprehensive evaluation of differential gene expression analysis methods for RNA-seq data
Franck Rapaport, Raya Khanin, Yupu Liang … (2013)
- Record: found
- Abstract: found
- Article: found
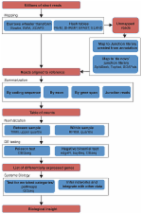
From RNA-seq reads to differential expression results
Alicia Oshlack, Mark Robinson, Matthew D. Young (2010)