- Record: found
- Abstract: found
- Article: found
YTHDC1-mediated augmentation of miR-30d in repressing pancreatic tumorigenesis via attenuation of RUNX1-induced transcriptional activation of Warburg effect
Read this article at
Abstract
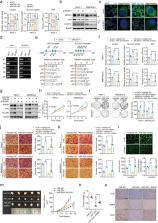
Pancreatic ductal adenocarcinoma (PDAC) is one of the most lethal human cancers. It thrives in a malnourished environment; however, little is known about the mechanisms by which PDAC cells actively promote aerobic glycolysis to maintain their metabolic needs. Gene Expression Omnibus (GEO) was used to identify differentially expressed miRNAs. The expression pattern of miR-30d in normal and PDAC tissues was studied by in situ hybridization. The role of miR-30d/RUNX1 in vitro and in vivo was evaluated by CCK8 assay and clonogenic formation as well as transwell experiment, subcutaneous xenograft model and liver metastasis model, respectively. Glucose uptake, ATP and lactate production were tested to study the regulatory effect of miR-30d/RUNX1 on aerobic glycolysis in PDAC cells. Quantitative real-time PCR, western blot, Chip assay, promoter luciferase activity, RIP, MeRIP, and RNA stability assay were used to explore the molecular mechanism of YTHDC1/miR-30d/RUNX1 in PDAC. Here, we discover that miR-30d expression was remarkably decreased in PDAC tissues and associated with good prognosis, contributed to the suppression of tumor growth and metastasis, and attenuation of Warburg effect. Mechanistically, the m 6A reader YTHDC1 facilitated the biogenesis of mature miR-30d via m 6A-mediated regulation of mRNA stability. Then, miR-30d inhibited aerobic glycolysis through regulating SLC2A1 and HK1 expression by directly targeting the transcription factor RUNX1, which bound to the promoters of the SLC2A1 and HK1 genes. Moreover, miR-30d was clinically inversely correlated with RUNX1, SLC2A1 and HK1, which function as adverse prognosis factors for overall survival in PDAC tissues. Overall, we demonstrated that miR-30d is a functional and clinical tumor-suppressive gene in PDAC. Our findings further uncover that miR-30d is a novel target for YTHDC1 through m 6A modification, and miR-30d represses pancreatic tumorigenesis via suppressing aerobic glycolysis.
Related collections
Most cited references42
- Record: found
- Abstract: found
- Article: not found
m6A-dependent regulation of messenger RNA stability
- Record: found
- Abstract: found
- Article: not found
Reading, writing and erasing mRNA methylation
- Record: found
- Abstract: found
- Article: found