- Record: found
- Abstract: found
- Article: found
Bimodal signatures of germline methylation are linked with gene expression plasticity in the coral Acropora millepora
Read this article at
Abstract
Background
In invertebrates, genes belonging to dynamically regulated functional categories appear to be less methylated than “housekeeping” genes, suggesting that DNA methylation may modulate gene expression plasticity. To date, however, experimental evidence to support this hypothesis across different natural habitats has been lacking.
Results
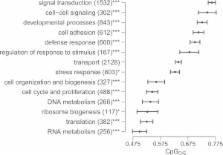
Gene expression profiles were generated from 30 pairs of genetically identical fragments of coral Acropora millepora reciprocally transplanted between distinct natural habitats for 3 months. Gene expression was analyzed in the context of normalized CpG content, a well-established signature of historical germline DNA methylation. Genes with weak methylation signatures were more likely to demonstrate differential expression based on both transplant environment and population of origin than genes with strong methylation signatures. Moreover, the magnitude of expression differences due to environment and population were greater for genes with weak methylation signatures.
Conclusions
Our results support a connection between differential germline methylation and gene expression flexibility across environments and populations. Studies of phylogenetically basal invertebrates such as corals will further elucidate the fundamental functional aspects of gene body methylation in Metazoa.
Related collections
Most cited references50
- Record: found
- Abstract: not found
- Book: not found
R: A language and environment for statistical computing
- Record: found
- Abstract: found
- Article: not found
Conserved Role of Intragenic DNA Methylation in Regulating Alternative Promoters
- Record: found
- Abstract: found
- Article: not found
Plant phenotypic plasticity in a changing climate.
Author and article information
Comments
Comment on this article
Similar content228
- A review of gorgonian coral species ( Cnidaria , Octocorallia , Alcyonacea ) held in the Santa Barbara Museum of Natural History research collection: focus on species from Scleraxonia , Holaxonia , Calcaxonia – Part II: Species of Holaxonia , families Gorgoniidae and PlexauridaeAuthors: Elizabeth Anne Horvath