- Record: found
- Abstract: found
- Article: found
Does co-transcriptional regulation of alternative splicing mediate plant stress responses?
Read this article at
Abstract
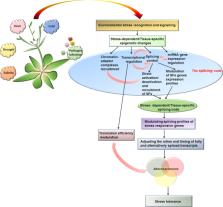
Plants display exquisite control over gene expression to elicit appropriate responses under normal and stress conditions. Alternative splicing (AS) of pre-mRNAs, a process that generates two or more transcripts from multi-exon genes, adds another layer of regulation to fine-tune condition-specific gene expression in animals and plants. However, exactly how plants control splice isoform ratios and the timing of this regulation in response to environmental signals remains elusive. In mammals, recent evidence indicate that epigenetic and epitranscriptome changes, such as DNA methylation, chromatin modifications and RNA methylation, regulate RNA polymerase II processivity, co-transcriptional splicing, and stability and translation efficiency of splice isoforms. In plants, the role of epigenetic modifications in regulating transcription rate and mRNA abundance under stress is beginning to emerge. However, the mechanisms by which epigenetic and epitranscriptomic modifications regulate AS and translation efficiency require further research. Dynamic changes in the chromatin landscape in response to stress may provide a scaffold around which gene expression, AS and translation are orchestrated. Finally, we discuss CRISPR/Cas-based strategies for engineering chromatin architecture to manipulate AS patterns (or splice isoforms levels) to obtain insight into the epigenetic regulation of AS.
Related collections
Most cited references110
- Record: found
- Abstract: found
- Article: not found
Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning.
- Record: found
- Abstract: found
- Article: not found
Alternative splicing: a pivotal step between eukaryotic transcription and translation.
- Record: found
- Abstract: found
- Article: not found
